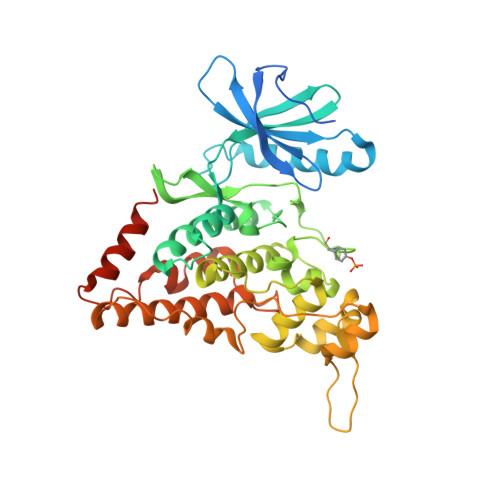
Abemaciclib is a potent inhibitor of DYRK1A and HIP kinases involved in transcriptional regulation.
Kaltheuner, I.H., Anand, K., Moecking, J., Duster, R., Wang, J., Gray, N.S., Geyer, M.(2021) Nat Commun 12: 6607-6607
- PubMed: 34785661 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-021-26935-z
- Primary Citation Related Structures:
7O7I, 7O7J, 7O7K - PubMed Abstract:
Homeodomain-interacting protein kinases (HIPKs) belong to the CMGC kinase family and are closely related to dual-specificity tyrosine phosphorylation-regulated kinases (DYRKs). HIPKs are regulators of various signaling pathways and involved in the pathology of cancer, chronic fibrosis, diabetes, and multiple neurodegenerative diseases. Here, we report the crystal structure of HIPK3 in its apo form at 2.5 Å resolution. Recombinant HIPKs and DYRK1A are auto-activated and phosphorylate the negative elongation factor SPT5, the transcription factor c-Myc, and the C-terminal domain of RNA polymerase II, suggesting a direct function in transcriptional regulation. Based on a database search, we identified abemaciclib, an FDA-approved Cdk4/Cdk6 inhibitor used for the treatment of metastatic breast cancer, as potent inhibitor of HIPK2, HIPK3, and DYRK1A. We determined the crystal structures of HIPK3 and DYRK1A bound to abemaciclib, showing a similar binding mode to the hinge region of the kinase as observed for Cdk6. Remarkably, DYRK1A is inhibited by abemaciclib to the same extent as Cdk4/Cdk6 in vitro, raising the question of whether targeting of DYRK1A contributes to the transcriptional inhibition and therapeutic activity of abemaciclib.
- Institute of Structural Biology, University of Bonn, Bonn, Germany.
Organizational Affiliation: