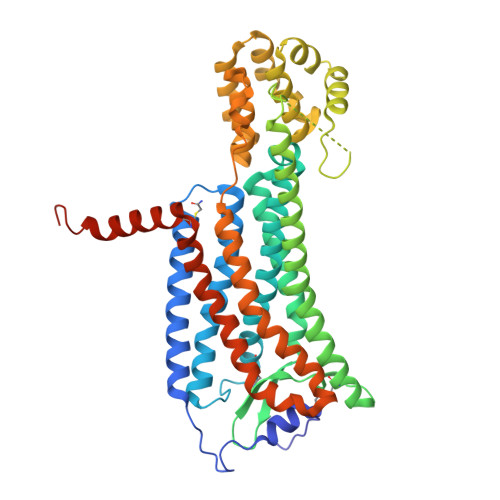
Molecular basis for lipid recognition by the prostaglandin D 2 receptor CRTH2.
Liu, H., Deepak, R.N.V.K., Shiriaeva, A., Gati, C., Batyuk, A., Hu, H., Weierstall, U., Liu, W., Wang, L., Cherezov, V., Fan, H., Zhang, C.(2021) Proc Natl Acad Sci U S A 118
- PubMed: 34341104 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.2102813118
- Primary Citation Related Structures:
7M8W - PubMed Abstract:
Prostaglandin D 2 (PGD 2 ) signals through the G protein-coupled receptor (GPCR) CRTH2 to mediate various inflammatory responses. CRTH2 is the only member of the prostanoid receptor family that is phylogenetically distant from others, implying a nonconserved mechanism of lipid action on CRTH2. Here, we report a crystal structure of human CRTH2 bound to a PGD 2 derivative, 15R-methyl-PGD 2 (15mPGD 2 ), by serial femtosecond crystallography. The structure revealed a "polar group in"-binding mode of 15mPGD 2 contrasting the "polar group out"-binding mode of PGE 2 in its receptor EP3. Structural comparison analysis suggested that these two lipid-binding modes, associated with distinct charge distributions of ligand-binding pockets, may apply to other lipid GPCRs. Molecular dynamics simulations together with mutagenesis studies also identified charged residues at the ligand entry port that function to capture lipid ligands of CRTH2 from the lipid bilayer. Together, our studies suggest critical roles of charge environment in lipid recognition by GPCRs.
- Department of Pharmacology and Chemical Biology, School of Medicine, University of Pittsburgh, Pittsburgh, PA 15261.
Organizational Affiliation: