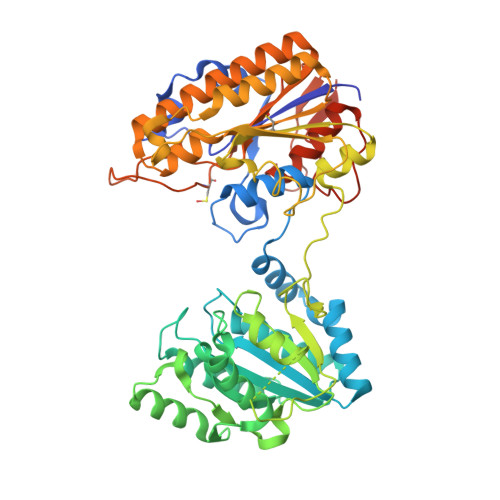
Structure-activity relationship of ipglycermide binding to phosphoglycerate mutases.
Wiedmann, M., Dranchak, P.K., Aitha, M., Queme, B., Collmus, C.D., Kashipathy, M.M., Kanter, L., Lamy, L., Rogers, J.M., Tao, D., Battaile, K.P., Rai, G., Lovell, S., Suga, H., Inglese, J.(2021) J Biological Chem 296: 100628-100628
- PubMed: 33812994 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jbc.2021.100628
- Primary Citation Related Structures:
7KNF, 7KNG - PubMed Abstract:
Catalysis of human phosphoglycerate mutase is dependent on a 2,3-bisphosphoglycerate cofactor (dPGM), whereas the nonhomologous isozyme in many parasitic species is cofactor independent (iPGM). This mechanistic and phylogenetic diversity offers an opportunity for selective pharmacologic targeting of glycolysis in disease-causing organisms. We previously discovered ipglycermide, a potent inhibitor of iPGM, from a large combinatorial cyclic peptide library. To fully delineate the ipglycermide pharmacophore, herein we construct a detailed structure-activity relationship using 280 substituted ipglycermide analogs. Binding affinities of these analogs to immobilized Caenorhabditis elegans iPGM, measured as fold enrichment relative to the index residue by deep sequencing of an mRNA display library, illuminated the significance of each amino acid to the pharmacophore. Using cocrystal structures and binding kinetics, we show that the high affinity of ipglycermide for iPGM orthologs, from Brugia malayi, Onchocerca volvulus, Dirofilaria immitis, and Escherichia coli, is achieved by a codependence between (1) the off-rate mediated by the macrocycle Cys14 thiolate coordination to an active-site Zn 2+ in the iPGM phosphatase domain and (2) shape complementarity surrounding the macrocyclic core at the phosphotransferase-phosphatase domain interface. Our results show that the high-affinity binding of ipglycermide to iPGMs freezes these structurally dynamic enzymes into an inactive, stable complex.
- Department of Chemistry, Graduate School of Sciences, The University of Tokyo, Tokyo, Japan.
Organizational Affiliation: