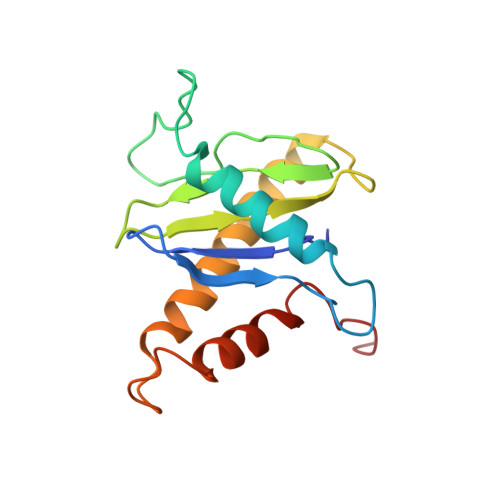
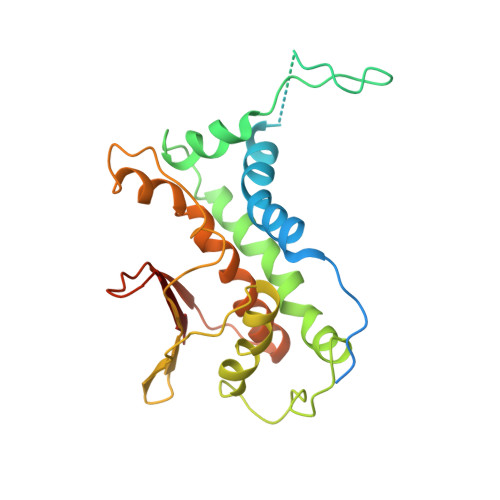
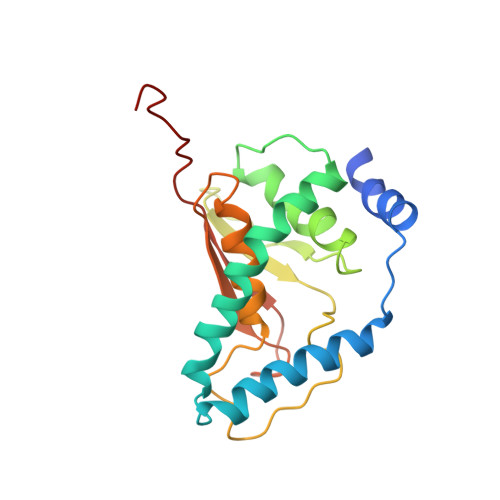
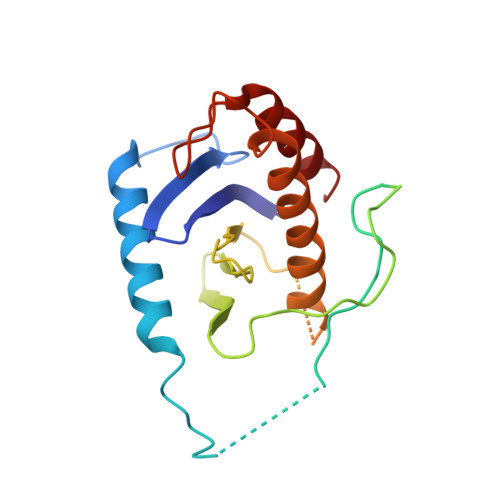
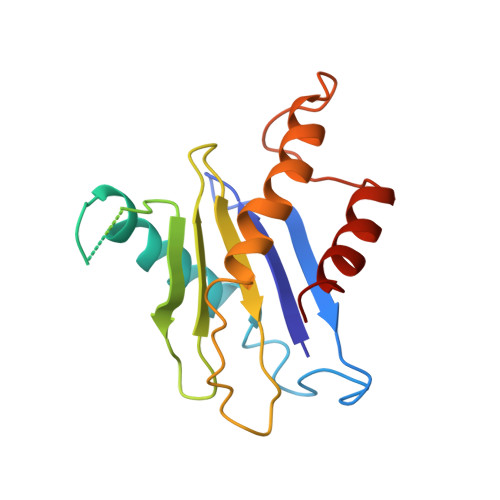
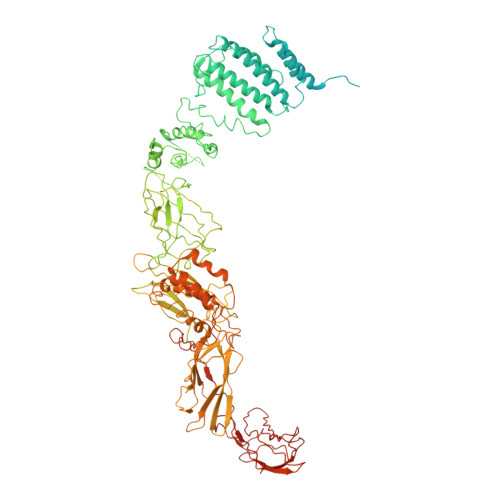
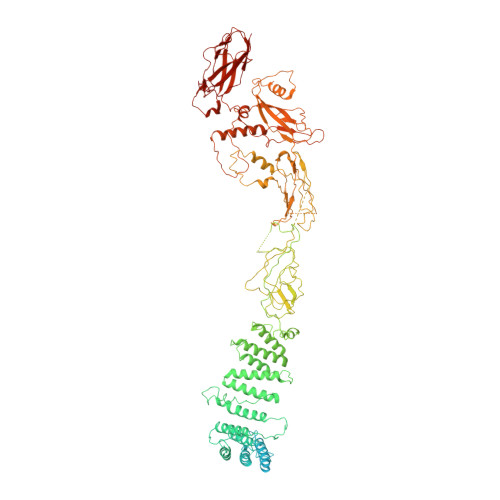
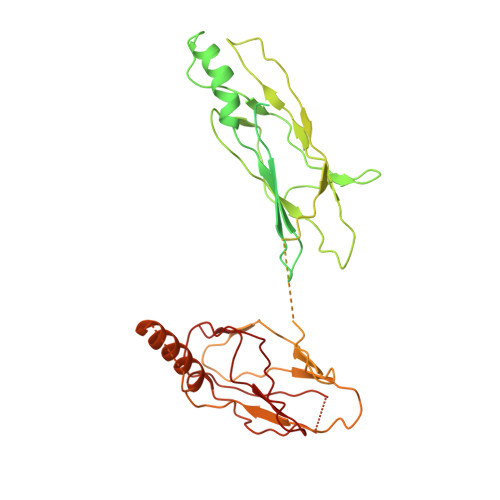
Structural basis for assembly of TRAPPII complex and specific activation of GTPase Ypt31/32.
Mi, C., Zhang, L., Huang, G., Shao, G., Yang, F., You, X., Dong, M.Q., Sun, S., Sui, S.F.(2022) Sci Adv 8: eabi5603-eabi5603
- PubMed: 35080977
- DOI: https://doi.org/10.1126/sciadv.abi5603
- Primary Citation Related Structures:
7E2C, 7E2D, 7E8S, 7E8T, 7E93, 7E94, 7EA3 - PubMed Abstract:
Transport protein particle (TRAPP) complexes belong to the multiprotein tethering complex and exist in three forms-core TRAPP/TRAPPI, TRAPPII, and TRAPPIII. TRAPPII activates GTPase Ypt31/Ypt32 as the guanine nucleotide exchange factor in the trans-Golgi network to determine the maturation of Golgi cisternae into post-Golgi carriers in yeast. Here, we present cryo-EM structures of yeast TRAPPII in apo and Ypt32-bound states. All the structures show a dimeric architecture assembled by two triangle-shaped monomers, while the monomer in the apo state exhibits both open and closed conformations, and the monomer in the Ypt32-bound form only captures the closed conformation. Located in the interior of the monomer, Ypt32 binds with both core TRAPP/TRAPPI and Trs120 via its nucleotide-binding domain and binds with Trs31 via its hypervariable domain. Combined with functional analysis, the structures provide insights into the assembly of TRAPPII and the mechanism of the specific activation of Ypt31/Ypt32 by TRAPPII.
- State Key Laboratory of Membrane Biology, Beijing Advanced Innovation Center for Structural Biology, Beijing Frontier Research Center for Biological Structure, School of Life Sciences, Tsinghua University, Beijing 100084, China.
Organizational Affiliation: