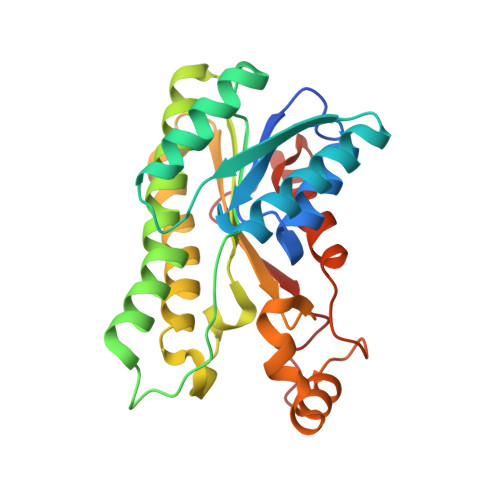
Crystal structure of l-rhamnose 1-dehydrogenase involved in the nonphosphorylative pathway of l-rhamnose metabolism in bacteria.
Yoshiwara, K., Watanabe, S., Watanabe, Y.(2021) FEBS Lett 595: 637-646
- PubMed: 33482017 Search on PubMed
- DOI: https://doi.org/10.1002/1873-3468.14046
- Primary Citation Related Structures:
7B81, 7DO5, 7DO6, 7DO7 - PubMed Abstract:
Several microorganisms can utilize l-rhamnose as a carbon and energy source through the nonphosphorylative metabolic pathway, in which l-rhamnose 1-dehydrogenase (RhaDH) catalyzes the NAD(P) + -dependent oxidization of l-rhamnose to l-rhamnono-1,4-lactone. We herein investigated the crystal structures of RhaDH from Azotobacter vinelandii in ligand-free, NAD + -bound, NADP + -bound, and l-rhamnose- and NAD + -bound forms at 1.9, 2.1, 2.4, and 1.6 Å resolution, respectively. The significant interactions with the 2'-phosphate group of NADP + , but not the 2'-hydroxyl group of NAD + , were consistent with a preference for NADP + over NAD + . The C5-OH and C6-methyl groups of l-rhamnose were recognized by specific residues of RhaDH through hydrogen bonds and hydrophobic contact, respectively, which contribute to the different substrate specificities from other aldose 1-dehydrogenases in the short-chain dehydrogenase/reductase superfamily.
- Faculty of Agriculture, Ehime University, Matsuyama, Japan.
Organizational Affiliation: