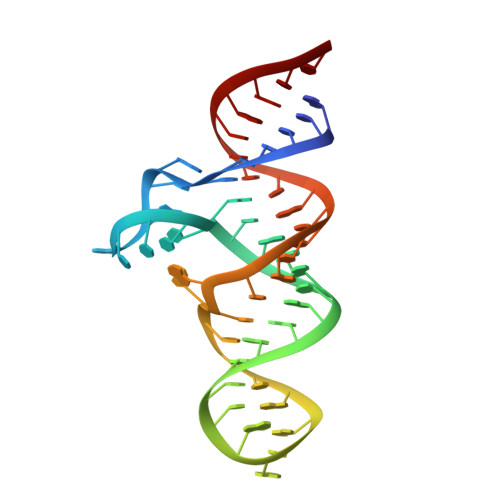
Structural distinctions between NAD+ riboswitch domains 1 and 2 determine differential folding and ligand binding.
Chen, H., Egger, M., Xu, X., Flemmich, L., Krasheninina, O., Sun, A., Micura, R., Ren, A.(2020) Nucleic Acids Res 48: 12394-12406
- PubMed: 33170270 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkaa1029
- Primary Citation Related Structures:
7D7V, 7D7W, 7D7X, 7D7Y, 7D7Z, 7D81, 7D82 - PubMed Abstract:
Riboswitches are important gene regulatory elements frequently encountered in bacterial mRNAs. The recently discovered nadA riboswitch contains two similar, tandemly arrayed aptamer domains, with the first domain possessing high affinity for nicotinamide adenine dinucleotide (NAD+). The second domain which comprises the ribosomal binding site in a putative regulatory helix, however, has withdrawn from detection of ligand-induced structural modulation thus far, and therefore, the identity of the cognate ligand and the regulation mechanism have remained unclear. Here, we report crystal structures of both riboswitch domains, each bound to NAD+. Furthermore, we demonstrate that ligand binding to domain 2 requires significantly higher concentrations of NAD+ (or ADP retaining analogs) compared to domain 1. Using a fluorescence spectroscopic approach, we further shed light on the structural features which are responsible for the different ligand affinities, and describe the Mg2+-dependent, distinct folding and pre-organization of their binding pockets. Finally, we speculate about possible scenarios for nadA RNA gene regulation as a putative two-concentration sensor module for a time-controlled signal that is primed and stalled by the gene regulation machinery at low ligand concentrations (domain 1), and finally triggers repression of translation as soon as high ligand concentrations are reached in the cell (domain 2).
- Life Sciences Institute, Zhejiang University, Hangzhou, Zhejiang 310058, China.
Organizational Affiliation: