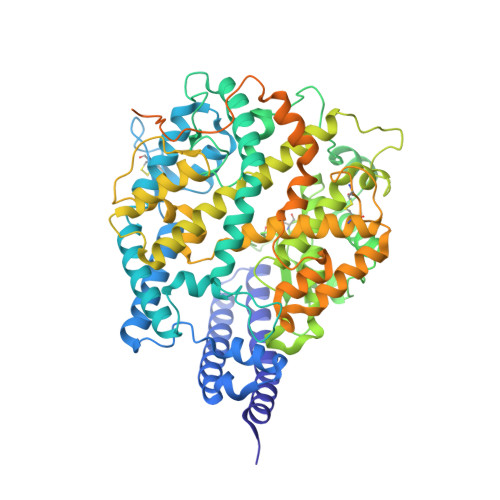
Broad host range of SARS-CoV-2 and the molecular basis for SARS-CoV-2 binding to cat ACE2.
Wu, L., Chen, Q., Liu, K., Wang, J., Han, P., Zhang, Y., Hu, Y., Meng, Y., Pan, X., Qiao, C., Tian, S., Du, P., Song, H., Shi, W., Qi, J., Wang, H.W., Yan, J., Gao, G.F., Wang, Q.(2020) Cell Discov 6: 68-68
- PubMed: 33020722 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41421-020-00210-9
- Primary Citation Related Structures:
7C8D - PubMed Abstract:
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), the causative agent of the recent pandemic COVID-19, is reported to have originated from bats, with its intermediate host unknown to date. Here, we screened 26 animal counterparts of the human ACE2 (hACE2), the receptor for SARS-CoV-2 and SARS-CoV, and found that the ACE2s from various species, including pets, domestic animals and multiple wild animals, could bind to SARS-CoV-2 receptor binding domain (RBD) and facilitate the transduction of SARS-CoV-2 pseudovirus. Comparing to SARS-CoV-2, SARS-CoV seems to have a slightly wider range in choosing its receptor. We further resolved the cryo-electron microscopy (cryo-EM) structure of the cat ACE2 (cACE2) in complex with the SARS-CoV-2 RBD at a resolution of 3 Å, revealing similar binding mode as hACE2 to the SARS-CoV-2 RBD. These results shed light on pursuing the intermediate host of SARS-CoV-2 and highlight the necessity of monitoring susceptible hosts to prevent further outbreaks.
- CAS Key Laboratory of Microbial Physiological and Metabolic Engineering, Institute of Microbiology, Chinese Academy of Sciences, Beijing, 100101 China.
Organizational Affiliation: