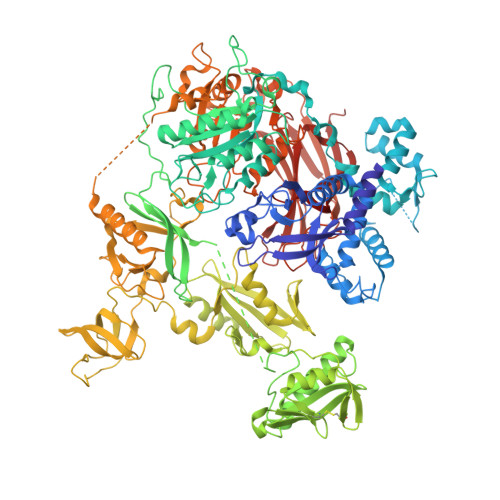
Characterization of the membrane interactions of phospholipase C gamma reveals key features of the active enzyme.
Le Huray, K.I.P., Bunney, T.D., Pinotsis, N., Kalli, A.C., Katan, M.(2022) Sci Adv 8: eabp9688-eabp9688
- PubMed: 35749497 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/sciadv.abp9688
- Primary Citation Related Structures:
7Z3J - PubMed Abstract:
PLCγ enzymes are autoinhibited in resting cells and form key components of intracellular signaling that are also linked to disease development. Insights into physiological and aberrant activation of PLCγ require understanding of an active, membrane-bound form, which can hydrolyze inositol-lipid substrates. Here, we demonstrate that PLCγ1 cannot bind membranes unless the autoinhibition is disrupted. Through extensive molecular dynamics simulations and experimental evidence, we characterize membrane binding by the catalytic core domains and reveal previously unknown sites of lipid interaction. The identified sites act in synergy, overlap with autoinhibitory interfaces, and are shown to be critical for the phospholipase activity in cells. This work provides direct evidence that PLCγ1 is inhibited through obstruction of its membrane-binding surfaces by the regulatory region and that activation must shift PLCγ1 to a conformation competent for membrane binding. Knowledge of the critical sites of membrane interaction extends the mechanistic framework for activation, dysregulation, and therapeutic intervention.
- Astbury Centre for Structural Molecular Biology, Faculty of Biological Sciences, University of Leeds, Leeds LS2 9JT, UK.
Organizational Affiliation: