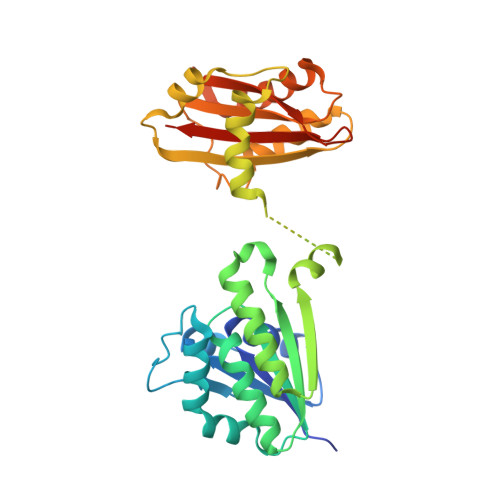
Dynamics of interdomain rotation facilitates FtsZ filament assembly.
Chakraborty, J., Poddar, S., Dutta, S., Bahulekar, V., Harne, S., Srinivasan, R., Gayathri, P.(2024) J Biological Chem 300: 107336-107336
- PubMed: 38718863 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jbc.2024.107336
- Primary Citation Related Structures:
7YOP, 7YSZ, 8GRW - PubMed Abstract:
FtsZ, the tubulin homolog essential for bacterial cell division, assembles as the Z-ring at the division site, and directs peptidoglycan synthesis by treadmilling. It is unclear how FtsZ achieves kinetic polarity that drives treadmilling. To obtain insights into fundamental features of FtsZ assembly dynamics independent of peptidoglycan synthesis, we carried out structural and biochemical characterization of FtsZ from the cell wall-less bacteria, Spiroplasma melliferum (SmFtsZ). Interestingly the structures of SmFtsZ, bound to GDP and GMPPNP respectively, were captured as domain swapped dimers. SmFtsZ was found to be a slower GTPase with a higher critical concentration (CC) compared to Escherichia coli FtsZ (EcFtsZ). In FtsZs, a conformational switch from R-state (close) to T-state (open) favors polymerization. We identified that Phe224, located at the interdomain cleft of SmFtsZ, is crucial for R- to T-state transition. SmFtsZ F224M exhibited higher GTPase activity and lower CC, whereas the corresponding EcFtsZ M225F resulted in cell division defects in E. coli. Our results demonstrate that relative rotation of the domains is a rate-limiting step of polymerization. Our structural analysis suggests that the rotation is plausibly triggered upon addition of a GTP-bound monomer to the filament through interaction of the preformed N-terminal domain (NTD). Hence, addition of monomers to the NTD-exposed end of filament is slower in comparison to the C-terminal domain (CTD) end, thus explaining kinetic polarity. In summary, the study highlights the importance of interdomain interactions and conformational changes in regulating FtsZ assembly dynamics.
- Biology Division, Indian Institute of Science Education and Research, Pune, India.
Organizational Affiliation: