Structural insight into ASH1L PHD finger recognizing methylated histone H3K4 and promoting cell growth in prostate cancer.
Yu, M., Jia, Y., Ma, Z., Ji, D., Wang, C., Liang, Y., Zhang, Q., Yi, H., Zeng, L.(2022) Front Oncol 12: 906807-906807
- PubMed: 36033518 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.3389/fonc.2022.906807
- Primary Citation Related Structures:
7Y0I - PubMed Abstract:
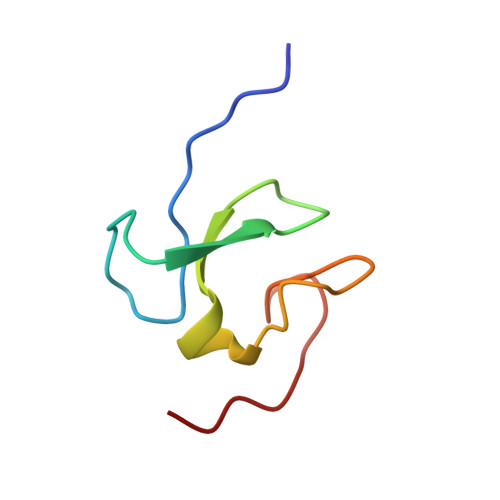
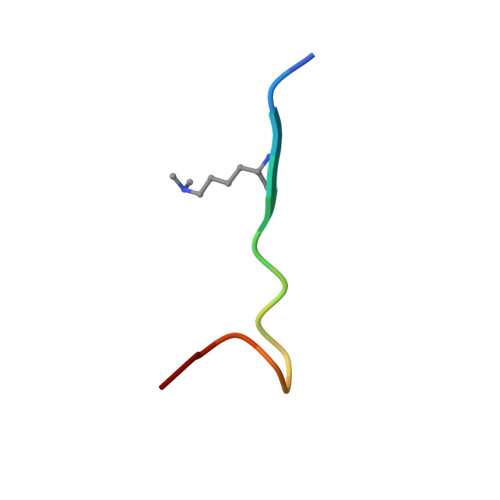
ASH1L is a member of the Trithorax-group protein and acts as a histone methyltransferase for gene transcription activation. It is known that ASH1L modulates H3K4me3 and H3K36me2/3 at its gene targets, but its specific mechanism of histone recognition is insufficiently understood. In this study, we found that the ASH1L plant homeodomain (PHD) finger interacts with mono-, di-, and trimethylated states of H3K4 peptides with comparable affinities, indicating that ASH1L PHD non-selectively binds to all three methylation states of H3K4. We solved nuclear magnetic resonance structures picturing the ASH1L PHD finger binding to the dimethylated H3K4 peptide and found that a narrow binding groove and residue composition in the methylated-lysine binding pocket restricts the necessary interaction with the dimethyl-ammonium moiety of K4. In addition, we found that the ASH1L protein is overexpressed in castrate-resistant prostate cancer (PCa) PC3 and DU145 cells in comparison to PCa LNCaP cells. The knockdown of ASH1L modulated gene expression and cellular pathways involved in apoptosis and cell cycle regulation and consequently induced cell cycle arrest, cell apoptosis, and reduced colony-forming abilities in PC3 and DU145 cells. The overexpression of the C-terminal core of ASH1L but not the PHD deletion mutant increased the overall H3K36me2 level but had no effect on the H3K4me2/3 level. Overall, our study identifies the ASH1L PHD finger as the first native reader that non-selectively recognizes the three methylation states of H3K4. Additionally, ASH1L is required for the deregulation of cell cycle and survival in PCas.
- Bethune Institute of Epigenetic Medicine, The First Hospital, Jilin University, Changchun, China.
Organizational Affiliation: