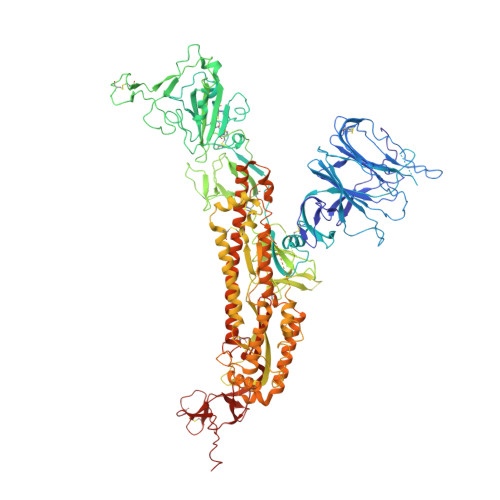
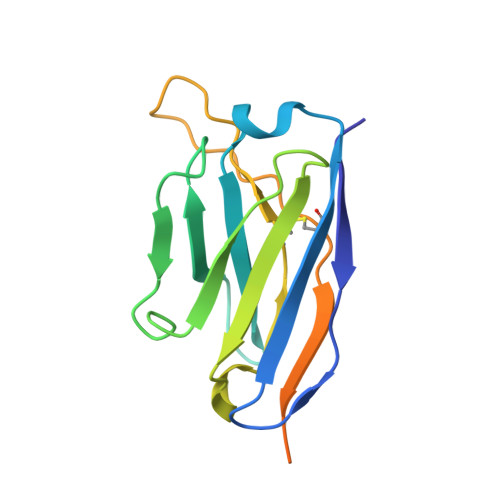
Identification of a nanobody able to catalyze the destruction of the spike-trimer of SARS-CoV-2.
Wang, K., Cao, D., Liu, L., Fan, X., Lin, Y., He, W., Zhai, Y., Xu, P., Yan, X., Wang, H., Zhang, X., Yang, P.(2025) Front Med 19: 493-506
- PubMed: 40317451 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1007/s11684-025-1128-4
- Primary Citation Related Structures:
7XY4 - PubMed Abstract:
Neutralizing antibodies have been designed to specifically target and bind to the receptor binding domain (RBD) of spike (S) protein to block severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) virus from attaching to angiotensin converting enzyme 2 (ACE2). This study reports a distinctive nanobody, designated as VHH21, that directly catalyzes the S-trimer into an irreversible transition state through postfusion conformational changes. Derived from camels immunized with multiple antigens, a set of nanobodies with high affinity for the S1 protein displays abilities to neutralize pseudovirion infections with a broad resistance to variants of concern of SARS-CoV-2, including SARS-CoV and BatRaTG13. Importantly, a super-resolution screening and analysis platform based on visual fluorescence probes was designed and applied to monitor single proteins and protein subunits. A spontaneously occurring dimeric form of VHH21 was obtained to rapidly destroy the S-trimer. Structural analysis via cryogenic electron microscopy revealed that VHH21 targets specific conserved epitopes on the S protein, distinct from the ACE2 binding site on the RBD, which destabilizes the fusion process. This research highlights the potential of VHH21 as an abzyme-like nanobody (nanoabzyme) possessing broad-spectrum binding capabilities and highly effective anti-viral properties and offers a promising strategy for combating coronavirus outbreaks.
- Key Laboratory of Epigenetic Regulation and Intervention, Institute of Biophysics, Chinese Academy of Sciences, Beijing, 100101, China.
Organizational Affiliation: