New mechanistic insight of cytochrome P450 PikC gained from site-specific mutagenesis by non-coding amino acids
Pan, Y.J., Li, G.B., Gao, X., Li, S.Y.(2023) Nat Commun
Experimental Data Snapshot
Starting Model: experimental
View more details
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Cytochrome P450 monooxygenase PikC | 444 | Streptomyces venezuelae | Mutation(s): 0 Gene Names: pikC, picK EC: 1.14.15.33 | 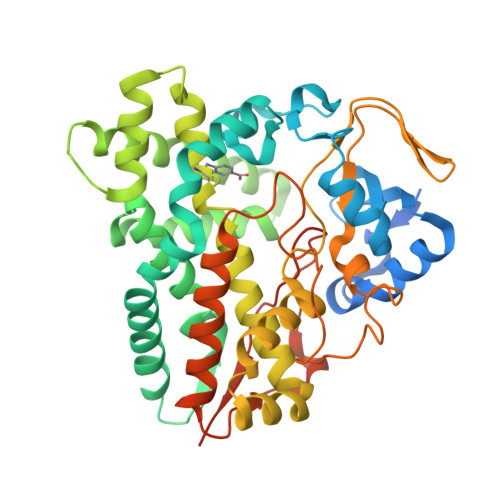 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | O87605 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 4 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| HEM Download:Ideal Coordinates CCD File | C [auth A], H [auth B] | PROTOPORPHYRIN IX CONTAINING FE C34 H32 Fe N4 O4 KABFMIBPWCXCRK-RGGAHWMASA-L |  | ||
| PXI (Subject of Investigation/LOI) Download:Ideal Coordinates CCD File | D [auth A], I [auth B] | 4-{[4-(DIMETHYLAMINO)-3-HYDROXY-6-METHYLTETRAHYDRO-2H-PYRAN-2-YL]OXY}-12-ETHYL-3,5,7,11-TETRAMETHYLOXACYCLODODEC-9-ENE-2,8-DIONE C25 H43 N O6 DZGHWPQKGWXOHD-OYZFACIPSA-N |  | ||
| PEG Download:Ideal Coordinates CCD File | F [auth A], G [auth A], J [auth B] | DI(HYDROXYETHYL)ETHER C4 H10 O3 MTHSVFCYNBDYFN-UHFFFAOYSA-N |  | ||
| GOL Download:Ideal Coordinates CCD File | E [auth A], K [auth B], L [auth B], M [auth B] | GLYCEROL C3 H8 O3 PEDCQBHIVMGVHV-UHFFFAOYSA-N |  | ||
| Modified Residues 1 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Type | Formula | 2D Diagram | Parent |
| 4AF Query on 4AF | A, B | L-PEPTIDE LINKING | C11 H13 N O3 |  | PHE |
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 59.823 | α = 90 |
| b = 108.903 | β = 90 |
| c = 152.63 | γ = 90 |
| Software Name | Purpose |
|---|---|
| PHENIX | refinement |
| HKL-3000 | data reduction |
| HKL-3000 | data scaling |
| PHENIX | phasing |
| Funding Organization | Location | Grant Number |
|---|---|---|
| National Natural Science Foundation of China (NSFC) | China | 32000039 |
| National Natural Science Foundation of China (NSFC) | China | 32025001 |
| National Natural Science Foundation of China (NSFC) | China | 31872729 |