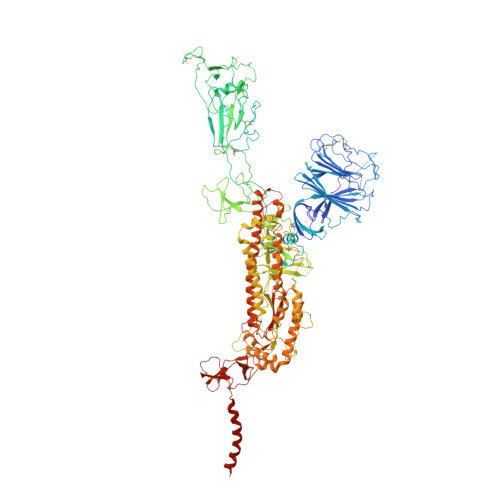
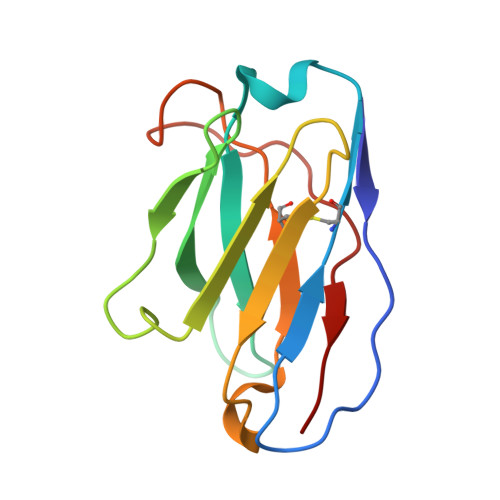
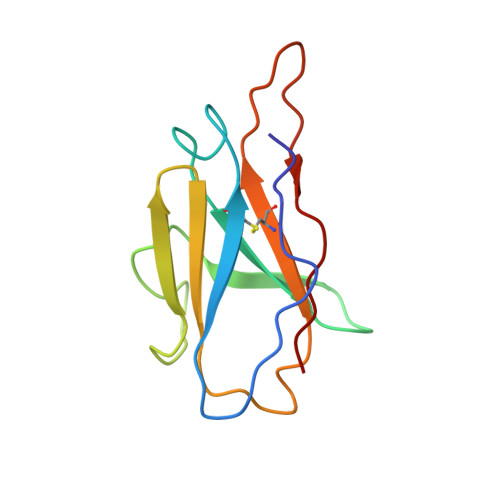
Selection and structural bases of potent broadly neutralizing antibodies from 3-dose vaccinees that are highly effective against diverse SARS-CoV-2 variants, including Omicron sublineages.
Wang, L., Fu, W., Bao, L., Jia, Z., Zhang, Y., Zhou, Y., Wu, W., Wu, J., Zhang, Q., Gao, Y., Wang, K., Wang, Q., Qin, C., Wang, X.(2022) Cell Res 32: 691-694
- PubMed: 35672388 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41422-022-00677-z
- Primary Citation Related Structures:
7WTF, 7WTG, 7WTH, 7WTI, 7WTJ, 7WTK - CAS Key Laboratory of Infection and Immunity, National Laboratory of Macromolecules, Institute of Biophysics, Chinese Academy of Sciences, Beijing, China.
Organizational Affiliation: