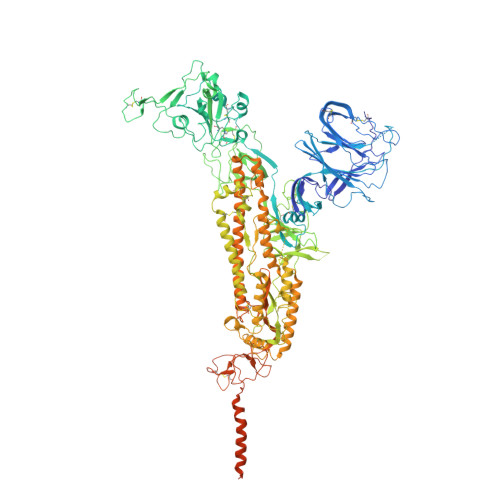
A broader neutralizing antibody against all the current VOCs and VOIs targets unique epitope of SARS-CoV-2 RBD.
Liu, S., Jia, Z., Nie, J., Liang, Z., Xie, J., Wang, L., Zhang, L., Wang, X., Wang, Y., Huang, W.(2022) Cell Discov 8: 81-81
- PubMed: 35977939 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41421-022-00443-w
- Primary Citation Related Structures:
7WT7, 7WT8, 7WT9 - Division of HIV/AIDS and Sex-transmitted Virus Vaccines, Institute for Biological Product Control, National Institutes for Food and Drug Control (NIFDC), Beijing, China.
Organizational Affiliation: