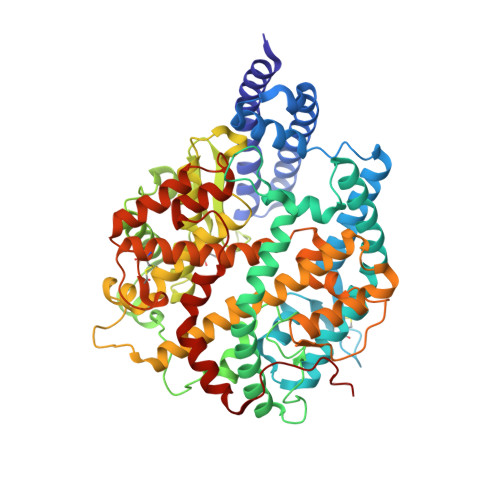
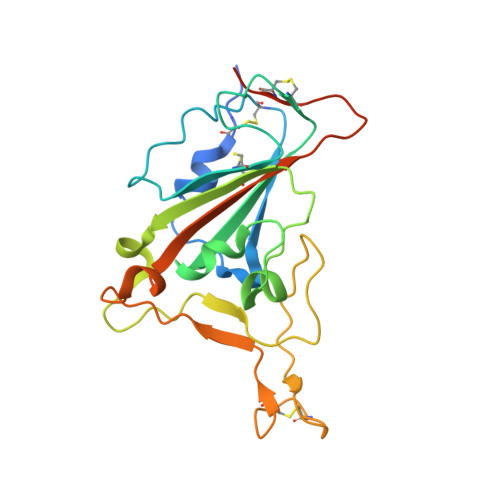
Receptor binding and complex structures of human ACE2 to spike RBD from omicron and delta SARS-CoV-2.
Han, P., Li, L., Liu, S., Wang, Q., Zhang, D., Xu, Z., Han, P., Li, X., Peng, Q., Su, C., Huang, B., Li, D., Zhang, R., Tian, M., Fu, L., Gao, Y., Zhao, X., Liu, K., Qi, J., Gao, G.F., Wang, P.(2022) Cell 185: 630-640.e10
- PubMed: 35093192 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.cell.2022.01.001
- Primary Citation Related Structures:
7WBL, 7WBP, 7WBQ - PubMed Abstract:
The coronavirus disease 2019 (COVID-19) pandemic continues worldwide with many variants arising, some of which are variants of concern (VOCs). A recent VOC, omicron (B.1.1.529), which obtains a large number of mutations in the receptor-binding domain (RBD) of the spike protein, has risen to intense scientific and public attention. Here, we studied the binding properties between the human receptor ACE2 (hACE2) and the VOC RBDs and resolved the crystal and cryoelectron microscopy structures of the omicron RBD-hACE2 complex as well as the crystal structure of the delta RBD-hACE2 complex. We found that, unlike alpha, beta, and gamma, omicron RBD binds to hACE2 at a similar affinity to that of the prototype RBD, which might be due to compensation of multiple mutations for both immune escape and transmissibility. The complex structures of omicron RBD-hACE2 and delta RBD-hACE2 reveal the structural basis of how RBD-specific mutations bind to hACE2.
- CAS Key Laboratory of Pathogen Microbiology and Immunology, Institute of Microbiology, Chinese Academy of Sciences, Beijing 100101, China; School of Medicine, Zhongda Hospital, Southeast University, NanJing 210009, China.
Organizational Affiliation: