Crystal Structure of HasAp with Co-5-octaethyloxaporphyrinium cation
Takiguchi, A., Sakakibara, E., Sugimoto, H., Shoji, O., Shinokubo, H.To be published.
Experimental Data Snapshot
Starting Model: experimental
View more details
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Heme acquisition protein HasAp | 184 | Pseudomonas aeruginosa PAO1 | Mutation(s): 0 Gene Names: hasAp, PA3407 | 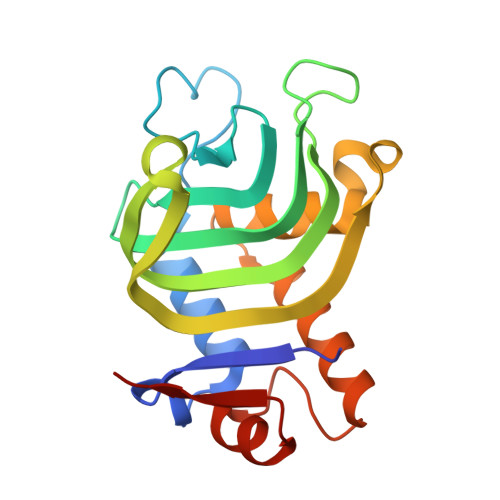 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | G3XD33 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 4 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| 7KI (Subject of Investigation/LOI) Download:Ideal Coordinates CCD File | C [auth A], H [auth B] | Co-5-octaethyloxaporphyrinium cation C35 H43 Co N4 O OPQNUXJQUPQEGE-SMMVCMDLSA-N |  | ||
| CIT Download:Ideal Coordinates CCD File | I [auth B] | CITRIC ACID C6 H8 O7 KRKNYBCHXYNGOX-UHFFFAOYSA-N |  | ||
| PEG Download:Ideal Coordinates CCD File | F [auth A], G [auth A] | DI(HYDROXYETHYL)ETHER C4 H10 O3 MTHSVFCYNBDYFN-UHFFFAOYSA-N |  | ||
| GOL Download:Ideal Coordinates CCD File | D [auth A], E [auth A] | GLYCEROL C3 H8 O3 PEDCQBHIVMGVHV-UHFFFAOYSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 153.84 | α = 90 |
| b = 153.84 | β = 90 |
| c = 37.751 | γ = 120 |
| Software Name | Purpose |
|---|---|
| XDS | data reduction |
| XSCALE | data scaling |
| REFMAC | refinement |
| PDB_EXTRACT | data extraction |
| REFMAC | phasing |
| Funding Organization | Location | Grant Number |
|---|---|---|
| Ministry of Education, Culture, Sports, Science and Technology (Japan) | Japan | JP17H01190 |
| Ministry of Education, Culture, Sports, Science and Technology (Japan) | Japan | JP19KK0138 |
| Ministry of Education, Culture, Sports, Science and Technology (Japan) | Japan | 17H05896 |
| Ministry of Education, Culture, Sports, Science and Technology (Japan) | Japan | 18H02396 |
| Ministry of Education, Culture, Sports, Science and Technology (Japan) | Japan | JP18H02084 |
| Japan Society for the Promotion of Science (JSPS) | Japan | JP20J11437 |
| Japan Society for the Promotion of Science (JSPS) | Japan | JP21J15614 |