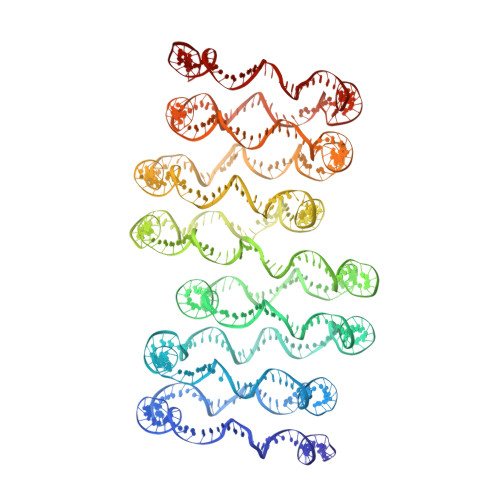
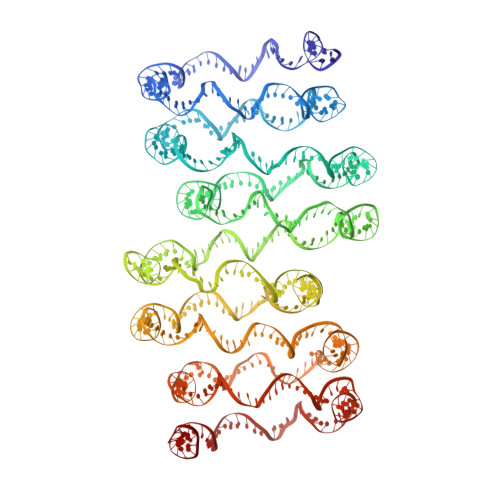
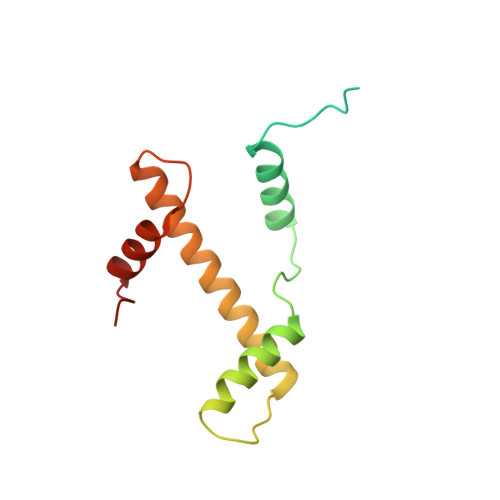
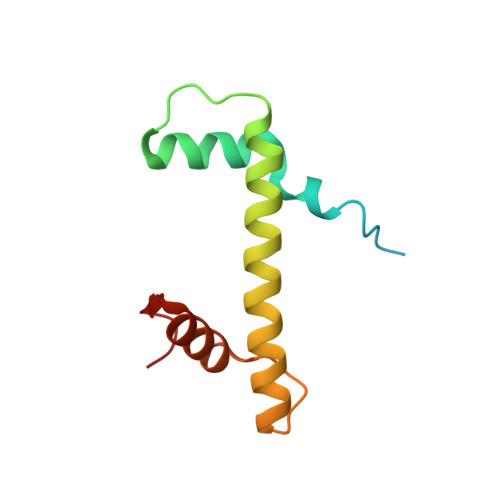
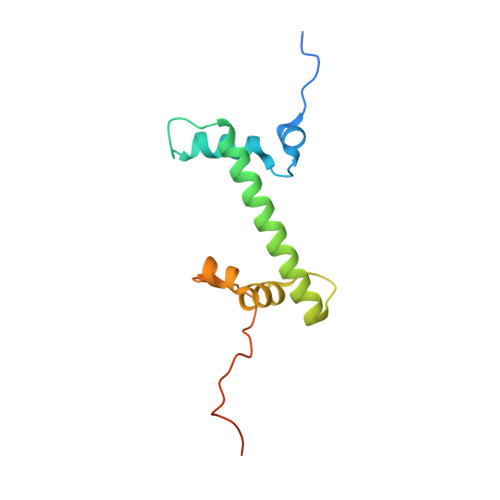
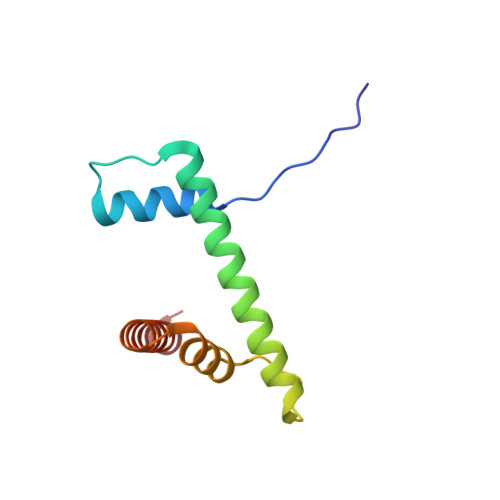
Columnar structure of human telomeric chromatin.
Soman, A., Wong, S.Y., Korolev, N., Surya, W., Lattmann, S., Vogirala, V.K., Chen, Q., Berezhnoy, N.V., van Noort, J., Rhodes, D., Nordenskiold, L.(2022) Nature 609: 1048-1055
- PubMed: 36104563 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41586-022-05236-5
- Primary Citation Related Structures:
7V90, 7V96, 7V9C, 7V9J, 7V9K, 7V9S, 7VA4 - PubMed Abstract:
Telomeres, the ends of eukaryotic chromosomes, play pivotal parts in ageing and cancer and are targets of DNA damage and the DNA damage response 1-5 . Little is known about the structure of telomeric chromatin at the molecular level. Here we used negative stain electron microscopy and single-molecule magnetic tweezers to characterize 3-kbp-long telomeric chromatin fibres. We also obtained the cryogenic electron microscopy structure of the condensed telomeric tetranucleosome and its dinucleosome unit. The structure displayed close stacking of nucleosomes with a columnar arrangement, and an unusually short nucleosome repeat length that comprised about 132 bp DNA wound in a continuous superhelix around histone octamers. This columnar structure is primarily stabilized by the H2A carboxy-terminal and histone amino-terminal tails in a synergistic manner. The columnar conformation results in exposure of the DNA helix, which may make it susceptible to both DNA damage and the DNA damage response. The conformation also exists in an alternative open state, in which one nucleosome is unstacked and flipped out, which exposes the acidic patch of the histone surface. The structural features revealed in this work suggest mechanisms by which protein factors involved in telomere maintenance can access telomeric chromatin in its compact form.
- School of Biological Sciences, Nanyang Technological University, Singapore, Singapore.
Organizational Affiliation: