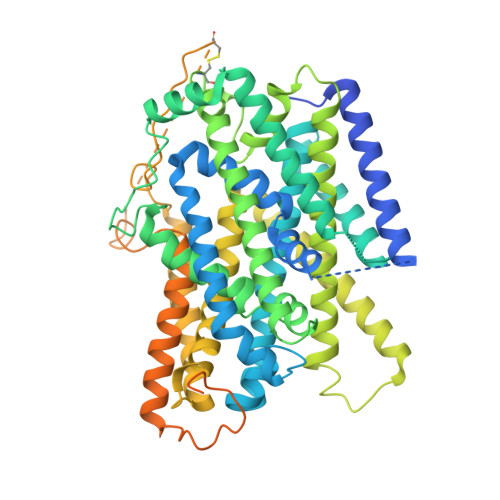
Structural insights into the mechanism of the sodium/iodide symporter.
Ravera, S., Nicola, J.P., Salazar-De Simone, G., Sigworth, F.J., Karakas, E., Amzel, L.M., Bianchet, M.A., Carrasco, N.(2022) Nature 612: 795-801
- PubMed: 36517601 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41586-022-05530-2
- Primary Citation Related Structures:
7UUY, 7UUZ, 7UV0 - PubMed Abstract:
The sodium/iodide symporter (NIS) is the essential plasma membrane protein that mediates active iodide (I - ) transport into the thyroid gland, the first step in the biosynthesis of the thyroid hormones-the master regulators of intermediary metabolism. NIS couples the inward translocation of I - against its electrochemical gradient to the inward transport of Na + down its electrochemical gradient 1,2 . For nearly 50 years before its molecular identification 3 , NIS was the molecule at the centre of the single most effective internal radiation cancer therapy: radioiodide ( 131 I - ) treatment for thyroid cancer 2 . Mutations in NIS cause congenital hypothyroidism, which must be treated immediately after birth to prevent stunted growth and cognitive deficiency 2 . Here we report three structures of rat NIS, determined by single-particle cryo-electron microscopy: one with no substrates bound; one with two Na + and one I - bound; and one with one Na + and the oxyanion perrhenate bound. Structural analyses, functional characterization and computational studies show the substrate-binding sites and key residues for transport activity. Our results yield insights into how NIS selects, couples and translocates anions-thereby establishing a framework for understanding NIS function-and how it transports different substrates with different stoichiometries and releases substrates from its substrate-binding cavity into the cytosol.
- Department of Molecular Physiology and Biophysics, Vanderbilt University, Nashville, TN, USA.
Organizational Affiliation: