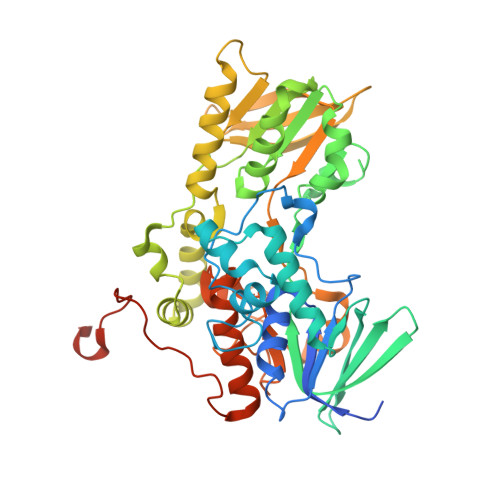
Kinetic and Structural Characterization of a Flavin-Dependent Putrescine N -Hydroxylase from Acinetobacter baumannii.
Lyons, N.S., Bogner, A.N., Tanner, J.J., Sobrado, P.(2022) Biochemistry 61: 2607-2620
- PubMed: 36314559 Search on PubMed
- DOI: https://doi.org/10.1021/acs.biochem.2c00493
- Primary Citation Related Structures:
7US3 - PubMed Abstract:
Acinetobacter baumannii is a Gram-negative opportunistic pathogen that causes nosocomial infections, especially among immunocompromised individuals. The rise of multidrug resistant strains of A. baumannii has limited the use of standard antibiotics, highlighting a need for new drugs that exploit novel mechanisms of pathogenicity. Disrupting iron acquisition by inhibiting the biosynthesis of iron-chelating molecules (siderophores) secreted by the pathogen is a potential strategy for developing new antibiotics. Here we investigated FbsI, an N -hydroxylating monooxygenase involved in the biosynthesis of fimsbactin A, the major siderophore produced by A. baumannii. FbsI was characterized using steady-state and transient-state kinetics, spectroscopy, X-ray crystallography, and small-angle X-ray scattering. FbsI was found to catalyze the N -hydroxylation of the aliphatic diamines putrescine and cadaverine. Maximum coupling of the reductive and oxidative half-reactions occurs with putrescine, suggesting it is the preferred ( in vivo ) substrate. FbsI uses both NADPH and NADH as the reducing cofactor with a slight preference for NADPH. The crystal structure of FbsI complexed with NADP + was determined at 2.2 Å resolution. The structure exhibits the protein fold characteristic of Class B flavin-dependent monooxygenases. FbsI is most similar in 3D structure to the cadaverine N -hydroxylases DesB and DfoA. Small-angle X-ray scattering shows that FbsI forms a tetramer in solution like the N -hydroxylating monooxygenases of the SidA/IucD/PvdA family. A model of putrescine docked into the active site provides insight into substrate recognition. A mechanism for the catalytic cycle is proposed where dehydration of the C4a-hydroxyflavin intermediate is partially rate-limiting, and the hydroxylated putrescine product is released before NADP + .
- Department of Biochemistry and Center for Drug Discovery, Virginia Tech, Blacksburg, Virginia 24061, United States.
Organizational Affiliation: