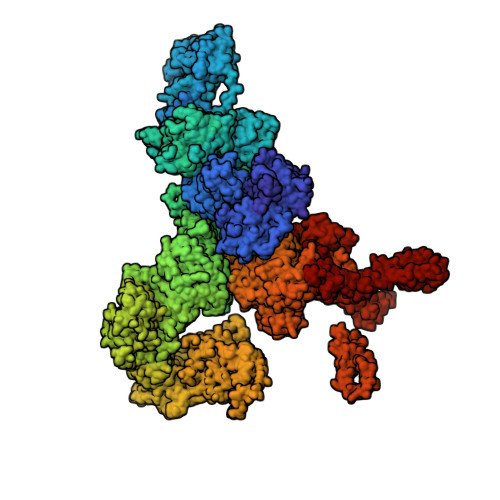
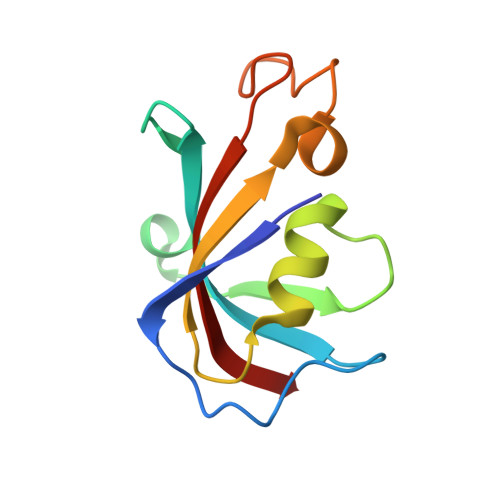
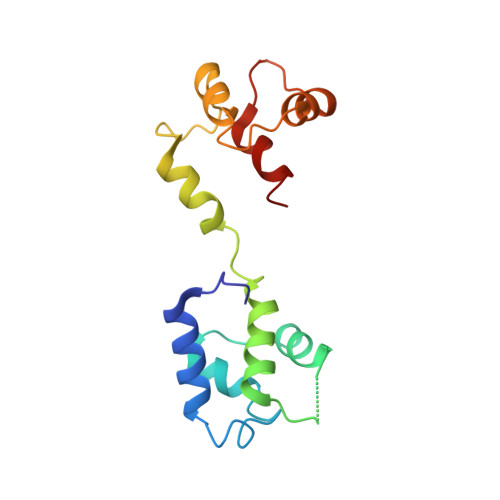
Structural analyses of human ryanodine receptor type 2 channels reveal the mechanisms for sudden cardiac death and treatment.
Miotto, M.C., Weninger, G., Dridi, H., Yuan, Q., Liu, Y., Wronska, A., Melville, Z., Sittenfeld, L., Reiken, S., Marks, A.R.(2022) Sci Adv 8: eabo1272-eabo1272
- PubMed: 35857850 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/sciadv.abo1272
- Primary Citation Related Structures:
7U9Q, 7U9R, 7U9T, 7U9X, 7U9Z, 7UA1, 7UA3, 7UA4, 7UA5, 7UA9 - PubMed Abstract:
Ryanodine receptor type 2 (RyR2) mutations have been linked to an inherited form of exercise-induced sudden cardiac death called catecholaminergic polymorphic ventricular tachycardia (CPVT). CPVT results from stress-induced sarcoplasmic reticular Ca 2+ leak via the mutant RyR2 channels during diastole. We present atomic models of human wild-type (WT) RyR2 and the CPVT mutant RyR2-R2474S determined by cryo-electron microscopy with overall resolutions in the range of 2.6 to 3.6 Å, and reaching local resolutions of 2.25 Å, unprecedented for RyR2 channels. Under nonactivating conditions, the RyR2-R2474S channel is in a "primed" state between the closed and open states of WT RyR2, rendering it more sensitive to activation that results in stress-induced Ca 2+ leak. The Rycal drug ARM210 binds to RyR2-R2474S, reverting the primed state toward the closed state. Together, these studies provide a mechanism for CPVT and for the therapeutic actions of ARM210.
- Department of Physiology and Cellular Biophysics, Columbia University Vagelos College of Physicians and Surgeons, New York, NY, USA.
Organizational Affiliation: