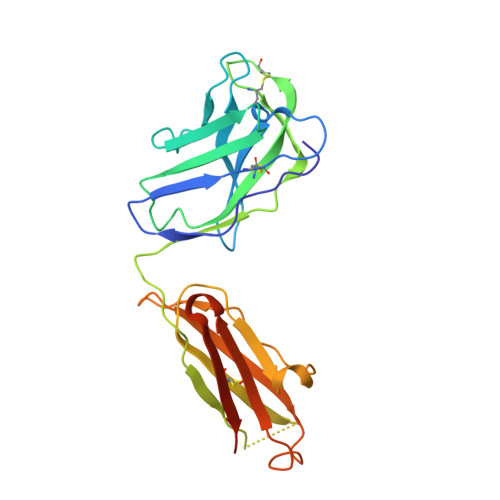
Computational identification of HCV neutralizing antibodies with a common HCDR3 disulfide bond motif in the antibody repertoires of infected individuals.
Bozhanova, N.G., Flyak, A.I., Brown, B.P., Ruiz, S.E., Salas, J., Rho, S., Bombardi, R.G., Myers, L., Soto, C., Bailey, J.R., Crowe Jr., J.E., Bjorkman, P.J., Meiler, J.(2022) Nat Commun 13: 3178-3178
- PubMed: 35676279 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-022-30865-9
- Primary Citation Related Structures:
7U0B, 7U0C - PubMed Abstract:
Despite recent success in hepatitis C virus (HCV) treatment using antivirals, an HCV vaccine is still needed to prevent reinfections in treated patients, to avert the emergence of drug-resistant strains, and to provide protection for people with no access to the antiviral therapeutics. The early production of broadly neutralizing antibodies (bNAbs) associates with HCV clearance. Several potent bNAbs bind a conserved HCV glycoprotein E2 epitope using an unusual heavy chain complementarity determining region 3 (HCDR3) containing an intra-loop disulfide bond. Isolation of additional structurally-homologous bNAbs would facilitate the recognition of key determinants of such bNAbs and guide rational vaccine design. Here we report the identification of new antibodies containing an HCDR3 disulfide bond motif using computational screening with the Rosetta software. Using the newly-discovered and already-known members of this antibody family, we review the required HCDR3 amino acid composition and propose determinants for the bent versus straight HCDR3 loop conformation observed in these antibodies.
- Department of Chemistry, Vanderbilt University, Nashville, TN, 37235, USA.
Organizational Affiliation: