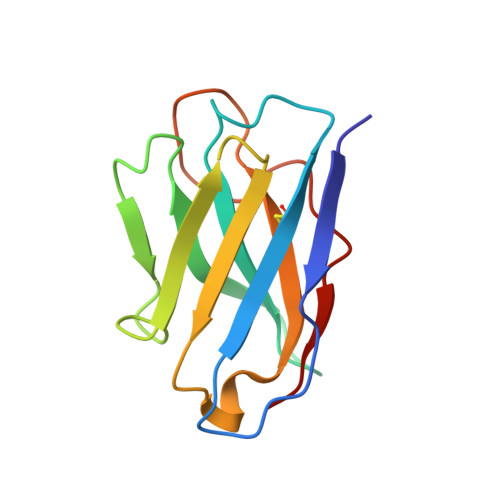
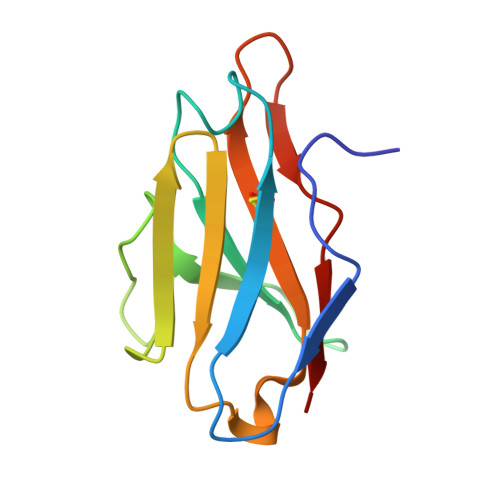
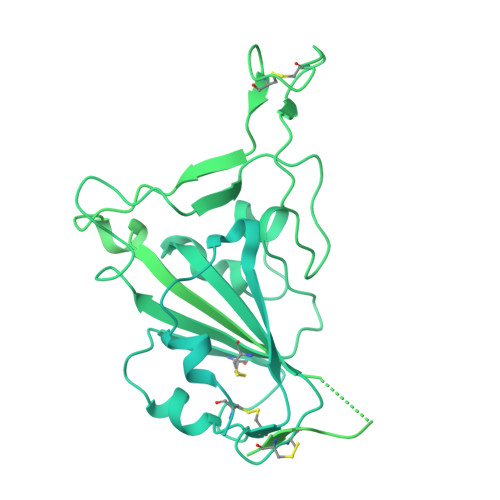
Antibody-mediated broad sarbecovirus neutralization through ACE2 molecular mimicry.
Park, Y.J., De Marco, A., Starr, T.N., Liu, Z., Pinto, D., Walls, A.C., Zatta, F., Zepeda, S.K., Bowen, J.E., Sprouse, K.R., Joshi, A., Giurdanella, M., Guarino, B., Noack, J., Abdelnabi, R., Foo, S.C., Rosen, L.E., Lempp, F.A., Benigni, F., Snell, G., Neyts, J., Whelan, S.P.J., Virgin, H.W., Bloom, J.D., Corti, D., Pizzuto, M.S., Veesler, D.(2022) Science 375: 449-454
- PubMed: 34990214 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/science.abm8143
- Primary Citation Related Structures:
7TAS, 7TAT - PubMed Abstract:
Understanding broadly neutralizing sarbecovirus antibody responses is key to developing countermeasures against SARS-CoV-2 variants and future zoonotic sarbecoviruses. We describe the isolation and characterization of a human monoclonal antibody, designated S2K146, that broadly neutralizes viruses belonging to SARS-CoV- and SARS-CoV-2-related sarbecovirus clades which use ACE2 as an entry receptor. Structural and functional studies show that most of the virus residues that directly bind S2K146 are also involved in binding to ACE2. This allows the antibody to potently inhibit receptor attachment. S2K146 protects against SARS-CoV-2 Beta challenge in hamsters and viral passaging experiments reveal a high barrier for emergence of escape mutants, making it a good candidate for clinical development. The conserved ACE2-binding residues present a site of vulnerability that might be leveraged for developing vaccines eliciting broad sarbecovirus immunity.
- Department of Biochemistry, University of Washington, Seattle, WA 98195, USA.
Organizational Affiliation: