Human antibodies to SARS-CoV-2 with a recurring YYDRxG motif retain binding and neutralization to variants of concern including Omicron.
Liu, H., Kaku, C.I., Song, G., Yuan, M., Andrabi, R., Burton, D.R., Walker, L.M., Wilson, I.A.(2022) Commun Biol 5: 766-766
- PubMed: 35906394 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s42003-022-03700-6
- Primary Citation Related Structures:
7T7B - PubMed Abstract:
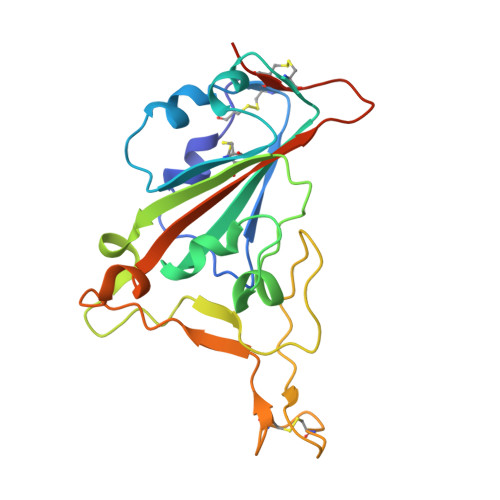
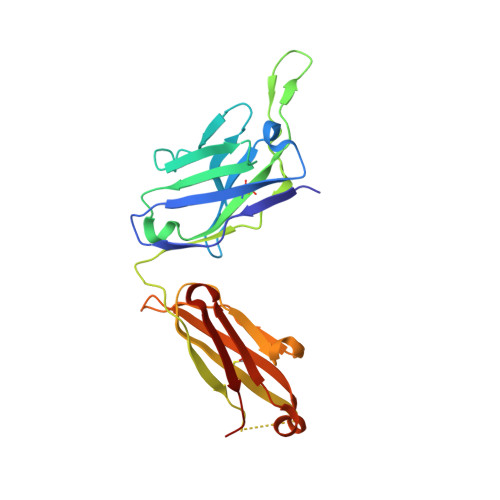
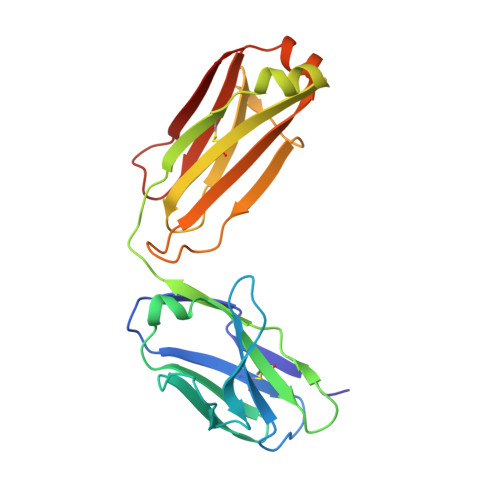
Studying the antibody response to SARS-CoV-2 informs on how the human immune system can respond to antigenic variants as well as other SARS-related viruses. Here, we structurally identified a YYDRxG motif encoded by IGHD3-22 in CDR H3 that facilitates antibody targeting to a functionally conserved epitope on the SARS-CoV-2 receptor binding domain. A computational search for a YYDRxG pattern in publicly available sequences uncovered 100 such antibodies, many of which can neutralize SARS-CoV-2 variants and SARS-CoV. Thus, the YYDRxG motif represents a common convergent solution for the human humoral immune system to target sarbecoviruses including the Omicron variant. These findings suggest an epitope-targeting strategy to identify potent and broadly neutralizing antibodies for design of pan-sarbecovirus vaccines and antibody therapeutics.
- Department of Integrative Structural and Computational Biology, The Scripps Research Institute, La Jolla, CA, USA.
Organizational Affiliation: