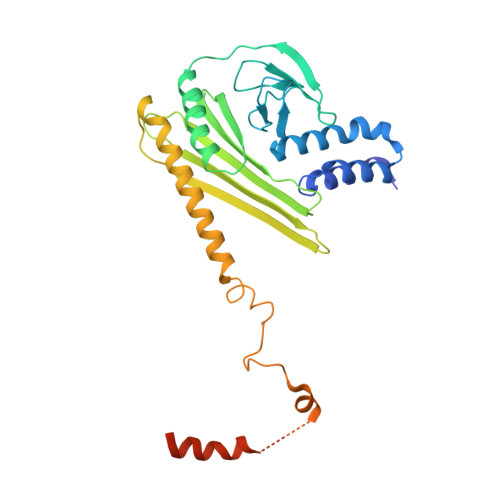
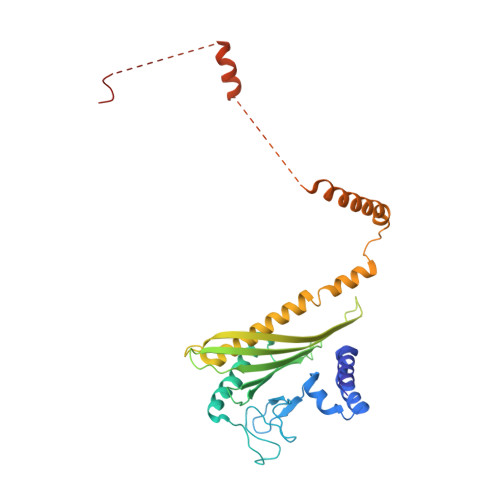
Transition State of Arp2/3 Complex Activation by Actin-Bound Dimeric Nucleation-Promoting Factor.
van Eeuwen, T., Boczkowska, M., Rebowski, G., Carman, P.J., Fregoso, F.E., Dominguez, R.(2023) Proc Natl Acad Sci U S A 120: e2306165120-e2306165120
- PubMed: 37549294 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.2306165120
- Primary Citation Related Structures:
7T5Q - PubMed Abstract:
Arp2/3 complex generates branched actin networks that drive fundamental processes such as cell motility and cytokinesis. The complex comprises seven proteins, including actin-related proteins (Arps) 2 and 3 and five scaffolding proteins (ArpC1-ArpC5) that mediate interactions with a pre-existing (mother) actin filament at the branch junction. Arp2/3 complex exists in two main conformations, inactive with the Arps interacting end-to-end and active with the Arps interacting side-by-side like subunits of the short-pitch helix of the actin filament. Several cofactors drive the transition toward the active state, including ATP binding to the Arps, WASP-family nucleation-promoting factors (NPFs), actin monomers, and binding of Arp2/3 complex to the mother filament. The precise contribution of each cofactor to activation is poorly understood. We report the 3.32-Å resolution cryo-electron microscopy structure of a transition state of Arp2/3 complex activation with bound constitutively dimeric NPF. Arp2/3 complex-binding region of the NPF N-WASP was fused C-terminally to the α and β subunits of the CapZ heterodimer. One arm of the NPF dimer binds Arp2 and the other binds actin and Arp3. The conformation of the complex is intermediate between those of inactive and active Arp2/3 complex. Arp2, Arp3, and actin also adopt intermediate conformations between monomeric (G-actin) and filamentous (F-actin) states, but only actin hydrolyzes ATP. In solution, the transition complex is kinetically shifted toward the short-pitch conformation and has higher affinity for F-actin than inactive Arp2/3 complex. The results reveal how all the activating cofactors contribute in a coordinated manner toward Arp2/3 complex activation.
- Biochemistry and Molecular Biophysics Graduate Group, Perelman School of Medicine, University of Pennsylvania, Philadelphia, PA 19104.
Organizational Affiliation: