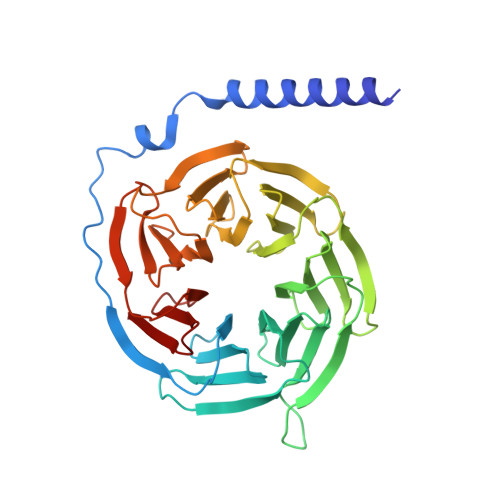
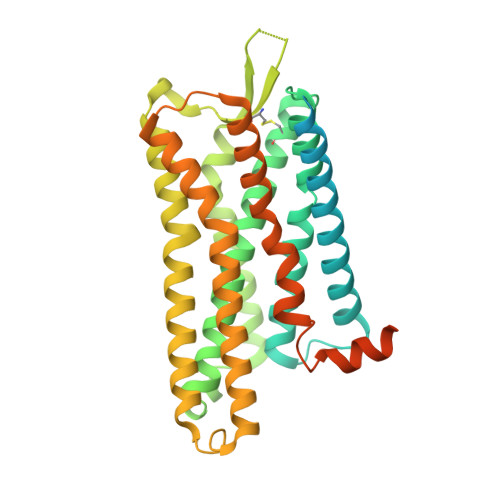
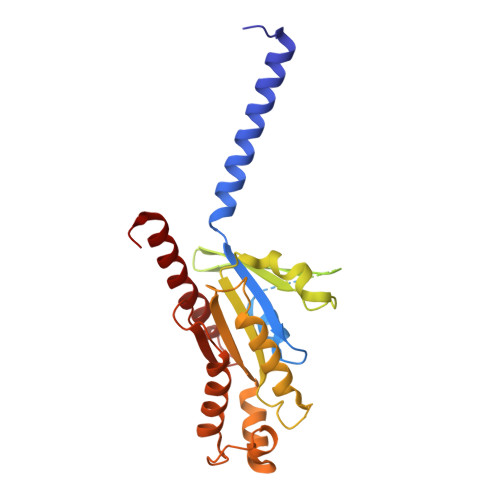
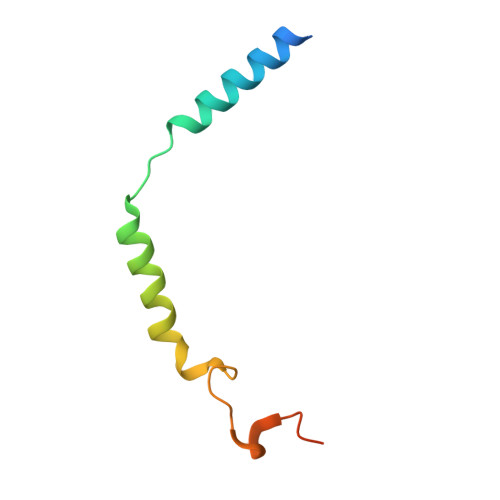
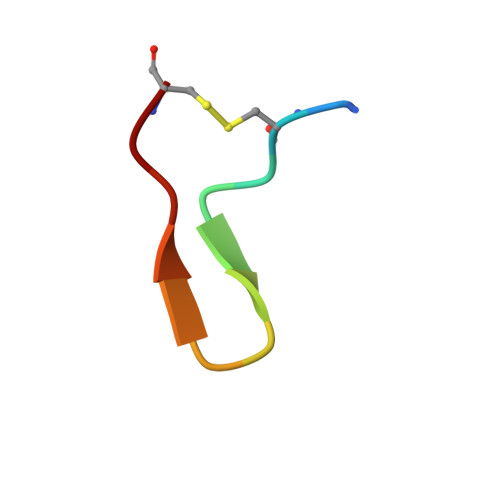
Plasticity in ligand recognition at somatostatin receptors.
Robertson, M.J., Meyerowitz, J.G., Panova, O., Borrelli, K., Skiniotis, G.(2022) Nat Struct Mol Biol 29: 210-217
- PubMed: 35210615 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41594-022-00727-5
- Primary Citation Related Structures:
7T10, 7T11 - PubMed Abstract:
Somatostatin is a signaling peptide that plays a pivotal role in physiologic processes relating to metabolism and growth through its actions at somatostatin receptors (SSTRs). Members of the SSTR subfamily, particularly SSTR2, are key drug targets for neuroendocrine neoplasms, with synthetic peptide agonists currently in clinical use. Here, we show the cryogenic-electron microscopy structures of active-state SSTR2 in complex with heterotrimeric G i3 and either the endogenous ligand SST14 or the FDA-approved drug octreotide. Complemented by biochemical assays and molecular dynamics simulations, these structures reveal key details of ligand recognition and receptor activation at SSTRs. We find that SSTR ligand recognition is highly diverse, as demonstrated by ligand-induced conformational changes in ECL2 and substantial sequence divergence across subtypes in extracellular regions. Despite this complexity, we rationalize several known sources of SSTR subtype selectivity and identify an additional interaction for specific binding. These results provide valuable insights for structure-based drug discovery at SSTRs.
- Department of Molecular and Cellular Physiology, Stanford University School of Medicine, Stanford, CA, USA.
Organizational Affiliation: