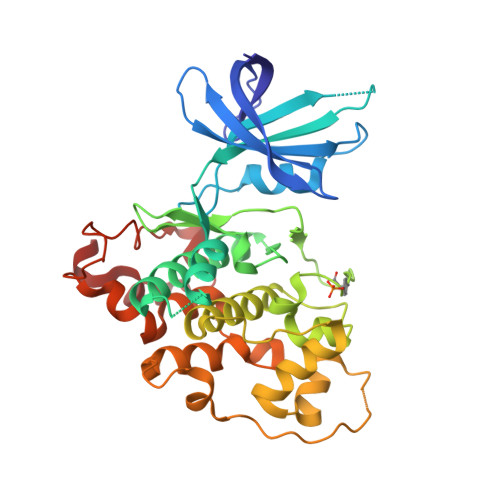
Elucidation of the GSK3 alpha Structure Informs the Design of Novel, Paralog-Selective Inhibitors.
Amaral, B., Capacci, A., Anderson, T., Tezer, C., Bajrami, B., Lulla, M., Lucas, B., Chodaparambil, J.V., Marcotte, D., Kumar, P.R., Murugan, P., Spilker, K., Cullivan, M., Wang, T., Peterson, A.C., Enyedy, I., Ma, B., Chen, T., Yousaf, Z., Calhoun, M., Golonzhka, O., Dillon, G.M., Koirala, S.(2023) ACS Chem Neurosci 14: 1080-1094
- PubMed: 36812145 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acschemneuro.2c00476
- Primary Citation Related Structures:
7SXF, 7SXG, 7SXH, 7SXJ - PubMed Abstract:
Glycogen synthase kinase 3 (GSK3) remains a therapeutic target of interest for diverse clinical indications. However, one hurdle in the development of small molecule GSK3 inhibitors has been safety concerns related to pan-inhibition of both GSK3 paralogs, leading to activation of the Wnt/β-catenin pathway and potential for aberrant cell proliferation. Development of GSK3α or GSK3β paralog-selective inhibitors that could offer an improved safety profile has been reported but further advancement has been hampered by the lack of structural information for GSK3α. Here we report for the first time the crystal structure for GSK3α, both in apo form and bound to a paralog-selective inhibitor. Taking advantage of this new structural information, we describe the design and in vitro testing of novel compounds with up to ∼37-fold selectivity for GSK3α over GSK3β with favorable drug-like properties. Furthermore, using chemoproteomics, we confirm that acute inhibition of GSK3α can lower tau phosphorylation at disease-relevant sites in vivo, with a high degree of selectivity over GSK3β and other kinases. Altogether, our studies advance prior efforts to develop GSK3 inhibitors by describing GSK3α structure and novel GSK3α inhibitors with improved selectivity, potency, and activity in disease-relevant systems.
- Departments of Research, Biogen, 225 Binney Street, Cambridge, Massachusetts 02142, United States.
Organizational Affiliation: