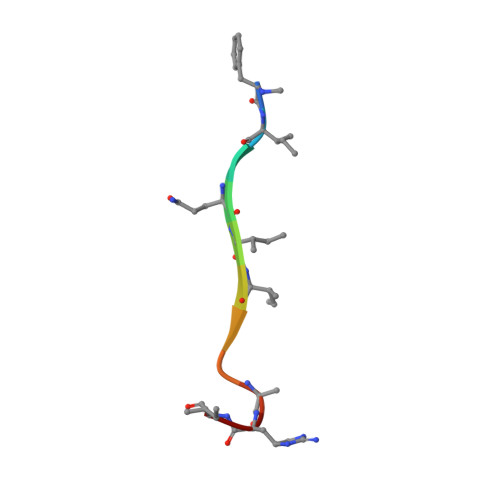
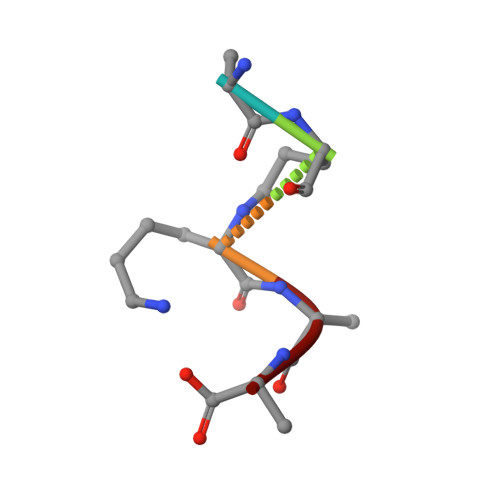
Teixobactin kills bacteria by a two-pronged attack on the cell envelope.
Shukla, R., Lavore, F., Maity, S., Derks, M.G.N., Jones, C.R., Vermeulen, B.J.A., Melcrova, A., Morris, M.A., Becker, L.M., Wang, X., Kumar, R., Medeiros-Silva, J., van Beekveld, R.A.M., Bonvin, A.M.J.J., Lorent, J.H., Lelli, M., Nowick, J.S., MacGillavry, H.D., Peoples, A.J., Spoering, A.L., Ling, L.L., Hughes, D.E., Roos, W.H., Breukink, E., Lewis, K., Weingarth, M.(2022) Nature 608: 390-396
- PubMed: 35922513 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41586-022-05019-y
- Primary Citation Related Structures:
7QGV - PubMed Abstract:
Antibiotics that use novel mechanisms are needed to combat antimicrobial resistance 1-3 . Teixobactin 4 represents a new class of antibiotics with a unique chemical scaffold and lack of detectable resistance. Teixobactin targets lipid II, a precursor of peptidoglycan 5 . Here we unravel the mechanism of teixobactin at the atomic level using a combination of solid-state NMR, microscopy, in vivo assays and molecular dynamics simulations. The unique enduracididine C-terminal headgroup of teixobactin specifically binds to the pyrophosphate-sugar moiety of lipid II, whereas the N terminus coordinates the pyrophosphate of another lipid II molecule. This configuration favours the formation of a β-sheet of teixobactins bound to the target, creating a supramolecular fibrillar structure. Specific binding to the conserved pyrophosphate-sugar moiety accounts for the lack of resistance to teixobactin 4 . The supramolecular structure compromises membrane integrity. Atomic force microscopy and molecular dynamics simulations show that the supramolecular structure displaces phospholipids, thinning the membrane. The long hydrophobic tails of lipid II concentrated within the supramolecular structure apparently contribute to membrane disruption. Teixobactin hijacks lipid II to help destroy the membrane. Known membrane-acting antibiotics also damage human cells, producing undesirable side effects. Teixobactin damages only membranes that contain lipid II, which is absent in eukaryotes, elegantly resolving the toxicity problem. The two-pronged action against cell wall synthesis and cytoplasmic membrane produces a highly effective compound targeting the bacterial cell envelope. Structural knowledge of the mechanism of teixobactin will enable the rational design of improved drug candidates.
- NMR Spectroscopy, Bijvoet Centre for Biomolecular Research, Department of Chemistry, Faculty of Science, Utrecht University, Utrecht, The Netherlands.
Organizational Affiliation: