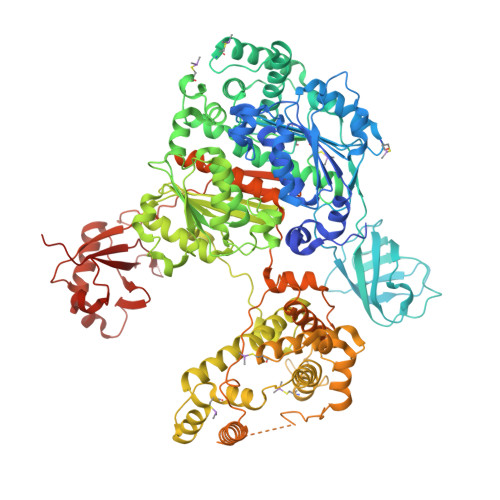
Structures of UBA6 explain its dual specificity for ubiquitin and FAT10.
Truongvan, N., Li, S., Misra, M., Kuhn, M., Schindelin, H.(2022) Nat Commun 13: 4789-4789
- PubMed: 35970836 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-022-32040-6
- Primary Citation Related Structures:
7PVN, 7PYV, 7ZH9 - PubMed Abstract:
The covalent modification of target proteins with ubiquitin or ubiquitin-like modifiers is initiated by E1 activating enzymes, which typically transfer a single modifier onto cognate conjugating enzymes. UBA6 is an unusual E1 since it activates two highly distinct modifiers, ubiquitin and FAT10. Here, we report crystal structures of UBA6 in complex with either ATP or FAT10. In the UBA6-FAT10 complex, the C-terminal domain of FAT10 binds to where ubiquitin resides in the UBA1-ubiquitin complex, however, a switch element ensures the alternate recruitment of either modifier. Simultaneously, the N-terminal domain of FAT10 interacts with the 3-helix bundle of UBA6. Site-directed mutagenesis identifies residues permitting the selective activation of either ubiquitin or FAT10. These results pave the way for studies investigating the activation of either modifier by UBA6 in physiological and pathophysiological settings.
- Institute of Structural Biology, Rudolf Virchow Center for Integrative and Translational Bioimaging, University of Würzburg, Josef-Schneider-Straße 2, 97080, Würzburg, Germany.
Organizational Affiliation: