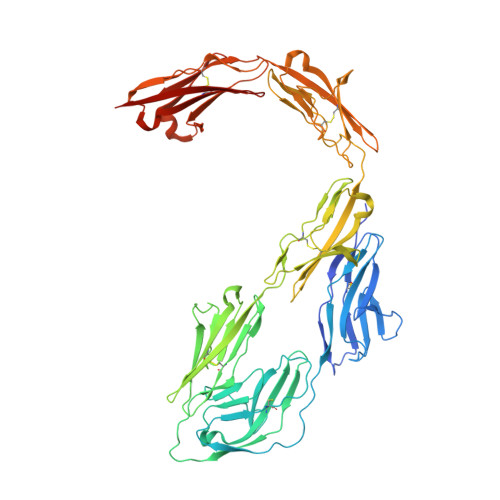
Structural insights into the contactin 1 - neurofascin 155 adhesion complex.
Chataigner, L.M.P., Gogou, C., den Boer, M.A., Frias, C.P., Thies-Weesie, D.M.E., Granneman, J.C.M., Heck, A.J.R., Meijer, D.H., Janssen, B.J.C.(2022) Nat Commun 13: 6607-6607
- PubMed: 36329006 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-022-34302-9
- Primary Citation Related Structures:
7OK5, 7OL2, 7OL4 - PubMed Abstract:
Cell-surface expressed contactin 1 and neurofascin 155 control wiring of the nervous system and interact across cells to form and maintain paranodal myelin-axon junctions. The molecular mechanism of contactin 1 - neurofascin 155 adhesion complex formation is unresolved. Crystallographic structures of complexed and individual contactin 1 and neurofascin 155 binding regions presented here, provide a rich picture of how competing and complementary interfaces, post-translational glycosylation, splice differences and structural plasticity enable formation of diverse adhesion sites. Structural, biophysical, and cell-clustering analysis reveal how conserved Ig1-2 interfaces form competing heterophilic contactin 1 - neurofascin 155 and homophilic neurofascin 155 complexes whereas contactin 1 forms low-affinity clusters through interfaces on Ig3-6. The structures explain how the heterophilic Ig1-Ig4 horseshoe's in the contactin 1 - neurofascin 155 complex define the 7.4 nm paranodal spacing and how the remaining six domains enable bridging of distinct intercellular distances.
- Structural Biochemistry, Bijvoet Center for Biomolecular Research, Faculty of Science, Utrecht University, Universiteitsweg 99, 3584 CG, Utrecht, The Netherlands.
Organizational Affiliation: