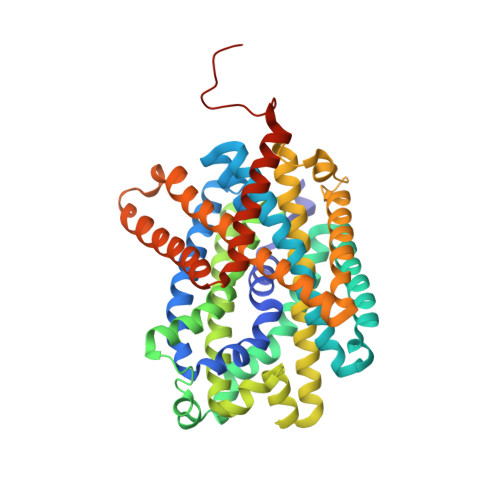
High-resolution structure of the amino acid transporter AdiC reveals insights into the role of water molecules and networks in oligomerization and substrate binding.
Ilgu, H., Jeckelmann, J.M., Kalbermatter, D., Ucurum, Z., Lemmin, T., Fotiadis, D.(2021) BMC Biol 19: 179-179
- PubMed: 34461897 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1186/s12915-021-01102-4
- Primary Citation Related Structures:
7O82 - PubMed Abstract:
The L-arginine/agmatine transporter AdiC is part of the arginine-dependent extreme acid resistance system of the bacterium Escherichia coli and its pathogenic varieties such as strain E. coli O157:H7. At the present time, there is a lack of knowledge concerning the role of water molecules and networks for the structure and function of AdiC, and solute transporters in general. The structure of the L-arginine/agmatine transporter AdiC was determined at 1.7 Å resolution by X-ray crystallography. This high resolution allowed for the identification of numerous water molecules buried in the structure. In combination with molecular dynamics (MD) simulations, we demonstrate that water molecules play an important role for stabilizing the protein and key residues, and act as placeholders for atoms of the AdiC substrates L-arginine and agmatine. MD simulations unveiled flexibility and restrained mobility of gating residues W202 and W293, respectively. Furthermore, a water-filled cavity was identified at the dimer interface of AdiC. The two monomers formed bridging interactions through water-mediated hydrogen bonds. The accessibility and presence of water molecules in this cavity was confirmed with MD simulations. Point mutations disrupting the interfacial water network validated the importance of water molecules for dimer stabilization. This work gives new insights into the role and importance of water molecules in the L-arginine/agmatine transporter AdiC for protein stabilization and substrate-binding site shaping and as placeholders of substrate atoms. Furthermore, and based on the observed flexibility and restrained mobility of gating residues, a mechanistic role of the gate flexibility in the transport cycle was proposed. Finally, we identified a water-filled cavity at the dimeric interface that contributes to the stability of the amino acid transporter oligomer.
- Institute of Biochemistry and Molecular Medicine, and Swiss National Centre of Competence in Research (NCCR) TransCure, University of Bern, CH-3012, Bern, Switzerland.
Organizational Affiliation: