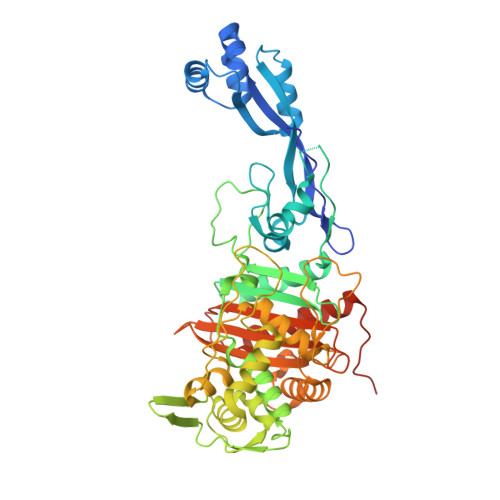
Structural analysis of the boronic acid beta-lactamase inhibitor vaborbactam binding to Pseudomonas aeruginosa penicillin-binding protein 3.
Kumar, V., Viviani, S.L., Ismail, J., Agarwal, S., Bonomo, R.A., van den Akker, F.(2021) PLoS One 16: e0258359-e0258359
- PubMed: 34653211 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0258359
- Primary Citation Related Structures:
7LY1 - PubMed Abstract:
Antimicrobial resistance (AMR) mediated by β-lactamases is the major and leading cause of resistance to penicillins and cephalosporins among Gram-negative bacteria. β-Lactamases, periplasmic enzymes that are widely distributed in the bacterial world, protect penicillin-binding proteins (PBPs), the major cell wall synthesizing enzymes, from inactivation by β-lactam antibiotics. Developing novel PBP inhibitors with a non-β-lactam scaffold could potentially evade this resistance mechanism. Based on the structural similarities between the evolutionary related serine β-lactamases and PBPs, we investigated whether the potent β-lactamase inhibitor, vaborbactam, could also form an acyl-enzyme complex with Pseudomonas aeruginosa PBP3. We found that this cyclic boronate, vaborbactam, inhibited PBP3 (IC50 of 262 μM), and its binding to PBP3 increased the protein thermal stability by about 2°C. Crystallographic analysis of the PBP3:vaborbactam complex reveals that vaborbactam forms a covalent bond with the catalytic S294. The amide moiety of vaborbactam hydrogen bonds with N351 and the backbone oxygen of T487. The carboxyl group of vaborbactam hydrogen bonds with T487, S485, and S349. The thiophene ring and cyclic boronate ring of vaborbactam form hydrophobic interactions, including with V333 and Y503. The active site of the vaborbactam-bound PBP3 harbors the often observed ligand-induced formation of the aromatic wall and hydrophobic bridge, yet the residues involved in this wall and bridge display much higher temperature factors compared to PBP3 structures bound to high-affinity β-lactams. These insights could form the basis for developing more potent novel cyclic boronate-based PBP inhibitors to inhibit these targets and overcome β-lactamases-mediated resistance mechanisms.
- Department of Biochemistry, Case Western Reserve University, Cleveland, Ohio, United States of America.
Organizational Affiliation: