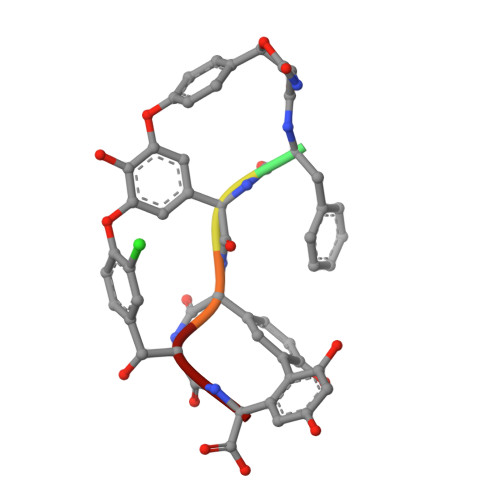
Potent and specific antibiotic combination therapy against Clostridioides difficile.
Chioti, V.T., McWhorter, K.L., Blue, T.C., Li, Y., Xu, F., Jeffrey, P.D., Davis, K.M., Seyedsayamdost, M.R.(2024) Nat Chem Biol 20: 924-933
- PubMed: 38942968 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41589-024-01651-z
- Primary Citation Related Structures:
7LKC, 7LTB - PubMed Abstract:
Keratinicyclins and keratinimicins are recently discovered glycopeptide antibiotics. Keratinimicins show broad-spectrum activity against Gram-positive bacteria, while keratinicyclins form a new chemotype by virtue of an unusual oxazolidinone moiety and exhibit specific antibiosis against Clostridioides difficile. Here we report the mechanism of action of keratinicyclin B (KCB). We find that steric constraints preclude KCB from binding peptidoglycan termini. Instead, KCB inhibits C. difficile growth by binding wall teichoic acids (WTAs) and interfering with cell wall remodeling. A computational model, guided by biochemical studies, provides an image of the interaction of KCB with C. difficile WTAs and shows that the same H-bonding framework used by glycopeptide antibiotics to bind peptidoglycan termini is used by KCB for interacting with WTAs. Analysis of KCB in combination with vancomycin (VAN) shows highly synergistic and specific antimicrobial activity, and that nanomolar combinations of the two drugs are sufficient for complete growth inhibition of C. difficile, while leaving common commensal strains unaffected.
- Department of Chemistry, Princeton University, Princeton, NJ, USA.
Organizational Affiliation: