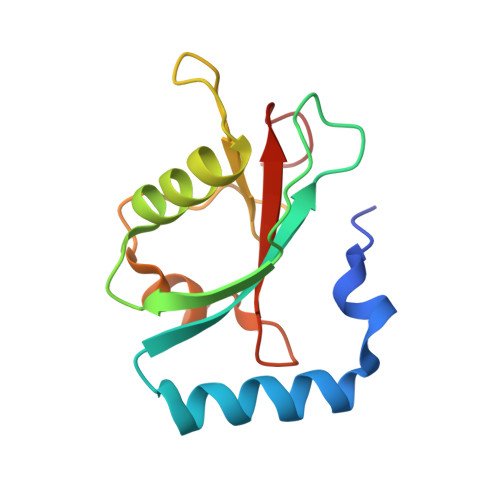
A new crystal form of GABARAPL2.
Scicluna, K., Dewson, G., Czabotar, P.E., Birkinshaw, R.W.(2021) Acta Crystallogr F Struct Biol Commun 77: 140-147
- PubMed: 33949974 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X21004489
- Primary Citation Related Structures:
7LK3 - PubMed Abstract:
The Atg8 protein family comprises the GABA type A receptor-associated proteins (GABARAPs) and microtubule-associated protein 1 light chains 3 (MAP1LC3s) that are essential mediators of autophagy. The LC3-interacting region (LIR) motifs of autophagy receptors and adaptors bind Atg8 proteins to promote autophagosome formation, cargo recruitment, and autophagosome closure and fusion to lysosomes. A crystal structure of human GABARAPL2 has been published [PDB entry 4co7; Ma et al. (2015), Biochemistry, 54, 5469-5479]. This was crystallized in space group P2 1 with a monoclinic angle of 90° and shows a pseudomerohedral twinning pathology. This article reports a new, untwinned GABARAPL2 crystal form, also in space group P2 1 , but with a 98° monoclinic angle. No major conformational differences were observed between the structures. In the structure described here, the C-terminal Phe117 binds into the LIR docking site (LDS) of a neighbouring molecule within the asymmetric unit, as observed in the previously reported structure. This crystal contact blocks the LDS for co-crystallization with ligands. Phe117 of GABARAPL2 is normally removed during biological processing by Atg4 family proteases. These data indicate that to establish interactions with the LIR, Phe117 should be removed to eliminate the crystal contact and liberate the LDS for co-crystallization with LIR peptides.
- Walter and Eliza Hall Institute of Medical Research, 1G Royal Parade, Parkville, VIC 3052, Australia.
Organizational Affiliation: