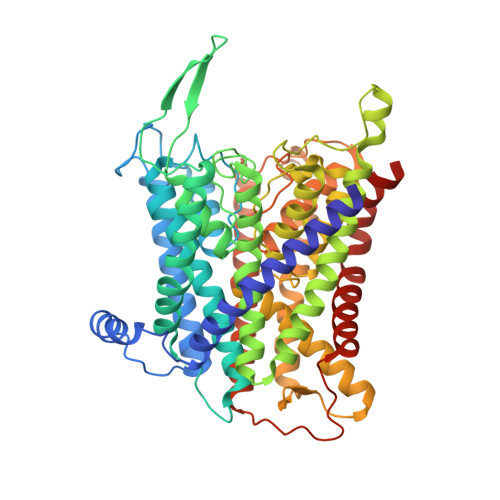
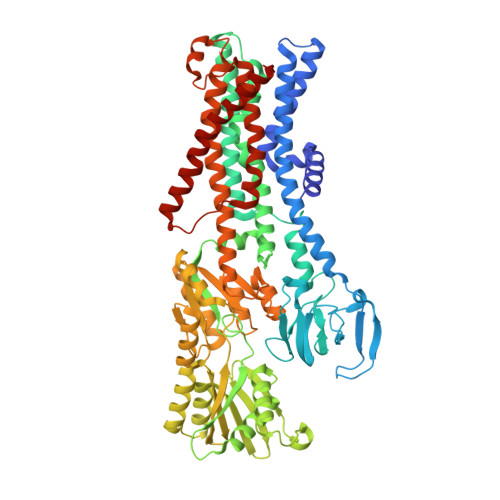
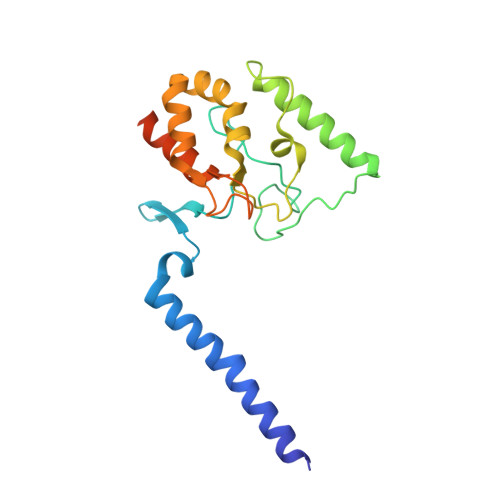
Structural basis for potassium transport in prokaryotes by KdpFABC.
Sweet, M.E., Larsen, C., Zhang, X., Schlame, M., Pedersen, B.P., Stokes, D.L.(2021) Proc Natl Acad Sci U S A 118
- PubMed: 34272288 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.2105195118
- Primary Citation Related Structures:
7BGY, 7BH1, 7BH2, 7LC3, 7LC6 - PubMed Abstract:
KdpFABC is an oligomeric K + transport complex in prokaryotes that maintains ionic homeostasis under stress conditions. The complex comprises a channel-like subunit (KdpA) from the superfamily of K + transporters and a pump-like subunit (KdpB) from the superfamily of P-type ATPases. Recent structural work has defined the architecture and generated contradictory hypotheses for the transport mechanism. Here, we use substrate analogs to stabilize four key intermediates in the reaction cycle and determine the corresponding structures by cryogenic electron microscopy. We find that KdpB undergoes conformational changes consistent with other representatives from the P-type superfamily, whereas KdpA, KdpC, and KdpF remain static. We observe a series of spherical densities that we assign as K + or water and which define a pathway for K + transport. This pathway runs through an intramembrane tunnel in KdpA and delivers ions to sites in the membrane domain of KdpB. Our structures suggest a mechanism where ATP hydrolysis is coupled to K + transfer between alternative sites in KdpB, ultimately reaching a low-affinity site where a water-filled pathway allows release of K + to the cytoplasm.
- Skirball Institute of Biomolecular Medicine, Department of Cell Biology, New York University Grossman School of Medicine, New York, NY 10016.
Organizational Affiliation: