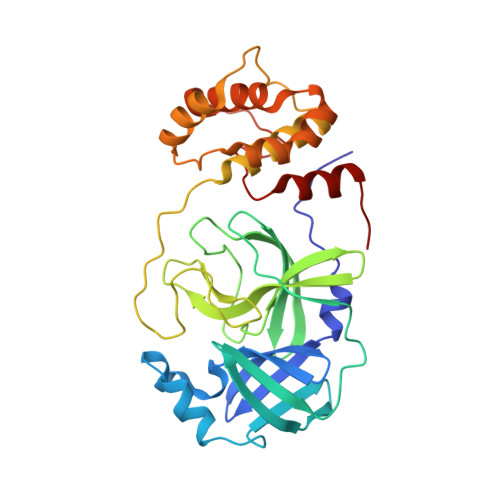
Pan-3C Protease Inhibitor Rupintrivir Binds SARS-CoV-2 Main Protease in a Unique Binding Mode.
Lockbaum, G.J., Henes, M., Lee, J.M., Timm, J., Nalivaika, E.A., Thompson, P.R., Kurt Yilmaz, N., Schiffer, C.A.(2021) Biochemistry 60: 2925-2931
- PubMed: 34506130 Search on PubMed
- DOI: https://doi.org/10.1021/acs.biochem.1c00414
- Primary Citation Related Structures:
7L8H, 7L8I, 7L8J - PubMed Abstract:
Rupintrivir targets the 3C cysteine proteases of the picornaviridae family, which includes rhinoviruses and enteroviruses that cause a range of human diseases. Despite being a pan-3C protease inhibitor, rupintrivir activity is extremely weak against the homologous 3C-like protease of SARS-CoV-2. In this study, the crystal structures of rupintrivir were determined bound to enterovirus 68 (EV68) 3C protease and the 3C-like main protease (M pro ) from SARS-CoV-2. While the EV68 3C protease-rupintrivir structure was similar to previously determined complexes with other picornavirus 3C proteases, rupintrivir bound in a unique conformation to the active site of SARS-CoV-2 M pro splitting the catalytic cysteine and histidine residues. This bifurcation of the catalytic dyad may provide a novel approach for inhibiting cysteine proteases.
- Department of Biochemistry and Molecular Pharmacology, University of Massachusetts Medical School, Worcester, Massachusetts 01605, United States.
Organizational Affiliation: