Genetic and structural basis for SARS-CoV-2 variant neutralization by a two-antibody cocktail.
Dong, J., Zost, S.J., Greaney, A.J., Starr, T.N., Dingens, A.S., Chen, E.C., Chen, R.E., Case, J.B., Sutton, R.E., Gilchuk, P., Rodriguez, J., Armstrong, E., Gainza, C., Nargi, R.S., Binshtein, E., Xie, X., Zhang, X., Shi, P.Y., Logue, J., Weston, S., McGrath, M.E., Frieman, M.B., Brady, T., Tuffy, K.M., Bright, H., Loo, Y.M., McTamney, P.M., Esser, M.T., Carnahan, R.H., Diamond, M.S., Bloom, J.D., Crowe Jr., J.E.(2021) Nat Microbiol 6: 1233-1244
- PubMed: 34548634 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41564-021-00972-2
- Primary Citation Related Structures:
7L7D, 7L7E - PubMed Abstract:
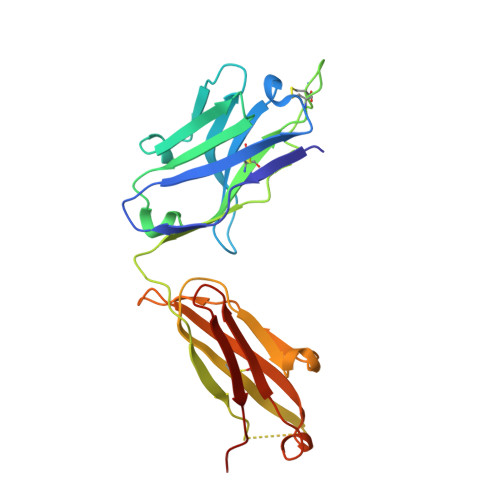
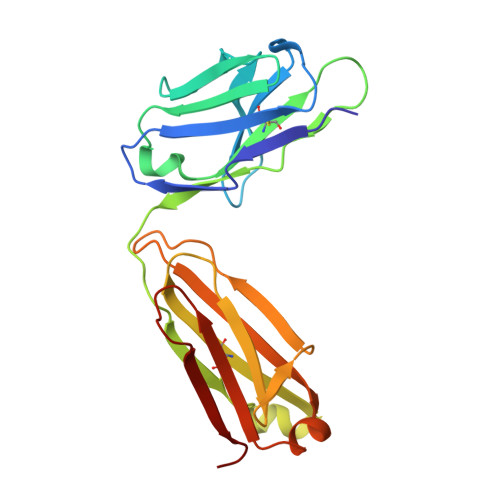
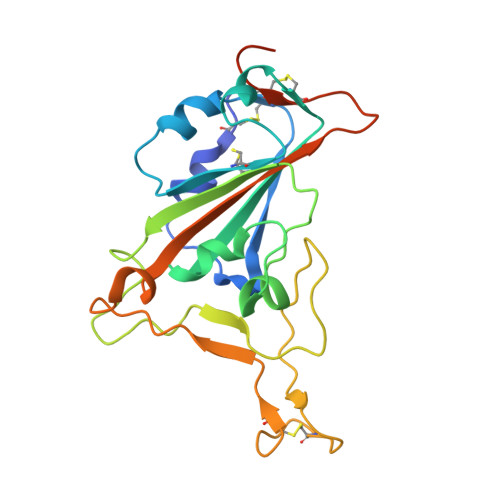
Understanding the molecular basis for immune recognition of SARS-CoV-2 spike glycoprotein antigenic sites will inform the development of improved therapeutics. We determined the structures of two human monoclonal antibodies-AZD8895 and AZD1061-which form the basis of the investigational antibody cocktail AZD7442, in complex with the receptor-binding domain (RBD) of SARS-CoV-2 to define the genetic and structural basis of neutralization. AZD8895 forms an 'aromatic cage' at the heavy/light chain interface using germ line-encoded residues in complementarity-determining regions (CDRs) 2 and 3 of the heavy chain and CDRs 1 and 3 of the light chain. These structural features explain why highly similar antibodies (public clonotypes) have been isolated from multiple individuals. AZD1061 has an unusually long LCDR1; the HCDR3 makes interactions with the opposite face of the RBD from that of AZD8895. Using deep mutational scanning and neutralization escape selection experiments, we comprehensively mapped the crucial binding residues of both antibodies and identified positions of concern with regards to virus escape from antibody-mediated neutralization. Both AZD8895 and AZD1061 have strong neutralizing activity against SARS-CoV-2 and variants of concern with antigenic substitutions in the RBD. We conclude that germ line-encoded antibody features enable recognition of the SARS-CoV-2 spike RBD and demonstrate the utility of the cocktail AZD7442 in neutralizing emerging variant viruses.
- Vanderbilt Vaccine Center, Vanderbilt University Medical Center, Nashville, TN, USA.
Organizational Affiliation: