LIG1 syndrome mutations remodel a cooperative network of ligand binding interactions to compromise ligation efficiency.
Jurkiw, T.J., Tumbale, P.P., Schellenberg, M.J., Cunningham-Rundles, C., Williams, R.S., O'Brien, P.J.(2021) Nucleic Acids Res 49: 1619-1630
- PubMed: 33444456 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkaa1297
- Primary Citation Related Structures:
7L34, 7L35 - PubMed Abstract:
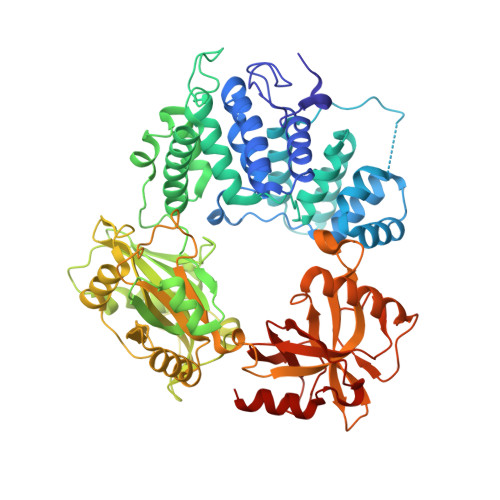
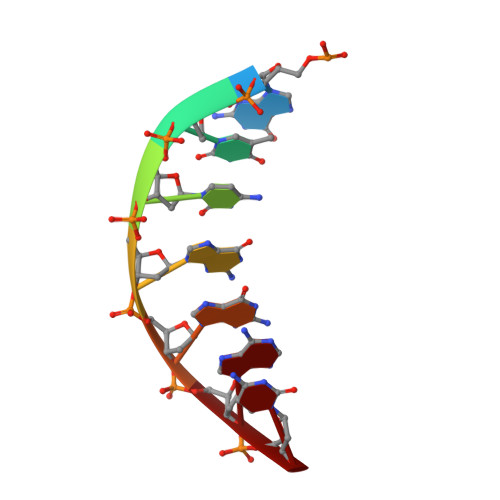
Human DNA ligase I (LIG1) is the main replicative ligase and it also seals DNA breaks to complete DNA repair and recombination pathways. Immune compromised patients harbor hypomorphic LIG1 alleles encoding substitutions of conserved arginine residues, R771W and R641L, that compromise LIG1 activity through poorly defined mechanisms. To understand the molecular basis of LIG1 syndrome mutations, we determined high resolution X-ray structures and performed systematic biochemical characterization of LIG1 mutants using steady-state and pre-steady state kinetic approaches. Our results unveil a cooperative network of plastic DNA-LIG1 interactions that connect DNA substrate engagement with productive binding of Mg2+ cofactors for catalysis. LIG1 syndrome mutations destabilize this network, compromising Mg2+ binding affinity, decreasing ligation efficiency, and leading to elevated abortive ligation that may underlie the disease pathology. These findings provide novel insights into the fundamental mechanism by which DNA ligases engage with a nicked DNA substrate, and they suggest that disease pathology of LIG1 syndrome could be modulated by Mg2+ levels.
- Department of Biological Chemistry, University of Michigan School of Medicine, Ann Arbor, MI, 48109, USA.
Organizational Affiliation: