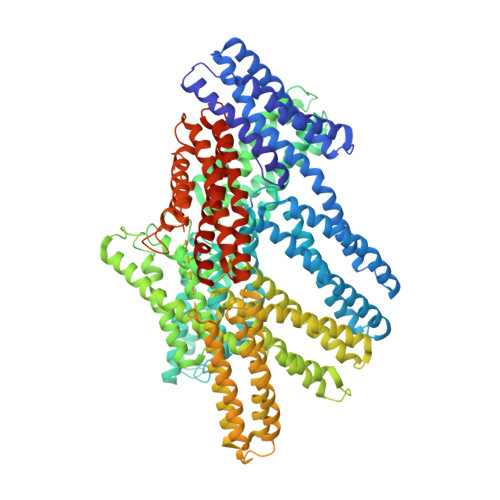
The Cryogenic Electron Microscopy Structure of the Cell Adhesion Regulator Metavinculin Reveals an Isoform-Specific Kinked Helix in Its Cytoskeleton Binding Domain.
Rangarajan, E.S., Izard, T.(2021) Int J Mol Sci 22
- PubMed: 33440717 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.3390/ijms22020645
- Primary Citation Related Structures:
7KTT, 7KTU, 7KTV, 7KTW - PubMed Abstract:
Vinculin and its heart-specific splice variant metavinculin are key regulators of cell adhesion processes. These membrane-bound cytoskeletal proteins regulate the cell shape by binding to several other proteins at cell-cell and cell-matrix junctions. Vinculin and metavinculin link integrin adhesion molecules to the filamentous actin network. Loss of both proteins prevents cell adhesion and cell spreading and reduces the formation of stress fibers, focal adhesions, or lamellipodia extensions. The binding of talin at cell-matrix junctions or of α-catenin at cell-cell junctions activates vinculin and metavinculin by releasing their autoinhibitory head-tail interaction. Once activated, vinculin and metavinculin bind F-actin via their five-helix bundle tail domains. Unlike vinculin, metavinculin has a 68-amino-acid insertion before the second α-helix of this five-helix F-actin-binding domain. Here, we present the full-length cryogenic electron microscopy structure of metavinculin that captures the dynamics of its individual domains and unveiled a hallmark structural feature, namely a kinked isoform-specific α-helix in its F-actin-binding domain. Our identified conformational landscape of metavinculin suggests a structural priming mechanism that is consistent with the cell adhesion functions of metavinculin in response to mechanical and cellular cues. Our findings expand our understanding of metavinculin function in the heart with implications for the etiologies of cardiomyopathies.
- Cell Adhesion Laboratory, Department of Integrative Structural and Computational Biology, The Scripps Research Institute, Jupiter, FL 33458, USA.
Organizational Affiliation: