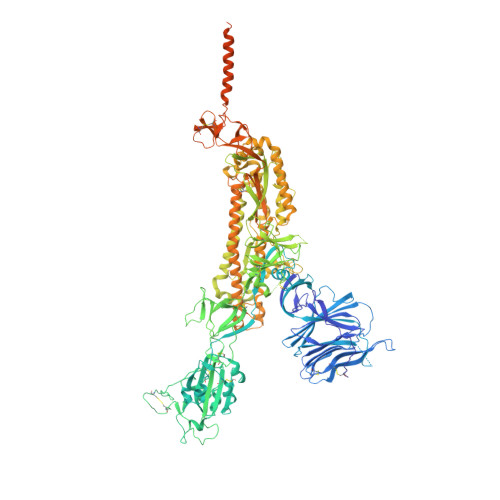
Structural impact on SARS-CoV-2 spike protein by D614G substitution.
Zhang, J., Cai, Y., Xiao, T., Lu, J., Peng, H., Sterling, S.M., Walsh Jr., R.M., Rits-Volloch, S., Zhu, H., Woosley, A.N., Yang, W., Sliz, P., Chen, B.(2021) Science 372: 525-530
- PubMed: 33727252 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/science.abf2303
- Primary Citation Related Structures:
7KRQ, 7KRR, 7KRS - PubMed Abstract:
Substitution for aspartic acid (D) by glycine (G) at position 614 in the spike (S) protein of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) appears to facilitate rapid viral spread. The G614 strain and its recent variants are now the dominant circulating forms. Here, we report cryo-electron microscopy structures of a full-length G614 S trimer, which adopts three distinct prefusion conformations that differ primarily by the position of one receptor-binding domain. A loop disordered in the D614 S trimer wedges between domains within a protomer in the G614 spike. This added interaction appears to prevent premature dissociation of the G614 trimer-effectively increasing the number of functional spikes and enhancing infectivity-and to modulate structural rearrangements for membrane fusion. These findings extend our understanding of viral entry and suggest an improved immunogen for vaccine development.
- Division of Molecular Medicine, Boston Children's Hospital, Boston, MA 02115, USA.
Organizational Affiliation: