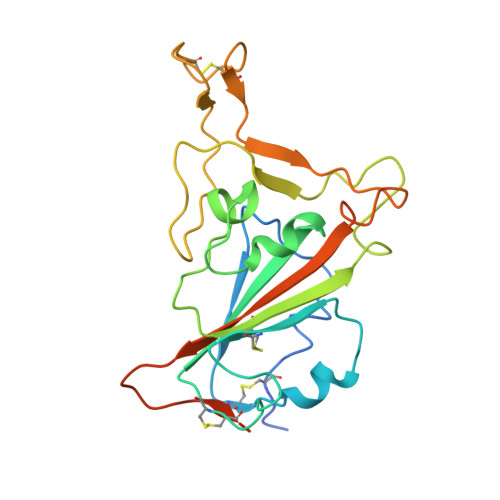
The development of Nanosota - 1 as anti-SARS-CoV-2 nanobody drug candidates.
Ye, G., Gallant, J., Zheng, J., Massey, C., Shi, K., Tai, W., Odle, A., Vickers, M., Shang, J., Wan, Y., Du, L., Aihara, H., Perlman, S., LeBeau, A., Li, F.(2021) Elife 10
- PubMed: 34338634 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.7554/eLife.64815
- Primary Citation Related Structures:
7KM5 - PubMed Abstract:
Combating the COVID-19 pandemic requires potent and low-cost therapeutics. We identified a series of single-domain antibodies (i.e., nanobody), Nanosota-1 , from a camelid nanobody phage display library. Structural data showed that Nanosota-1 bound to the oft-hidden receptor-binding domain (RBD) of SARS-CoV-2 spike protein, blocking viral receptor angiotensin-converting enzyme 2 (ACE2). The lead drug candidate possessing an Fc tag ( Nanosota-1C-Fc ) bound to SARS-CoV-2 RBD ~3000 times more tightly than ACE2 did and inhibited SARS-CoV-2 pseudovirus ~160 times more efficiently than ACE2 did. Administered at a single dose, Nanosota-1C-Fc demonstrated preventive and therapeutic efficacy against live SARS-CoV-2 infection in both hamster and mouse models. Unlike conventional antibodies, Nanosota-1C-Fc was produced at high yields in bacteria and had exceptional thermostability. Pharmacokinetic analysis of Nanosota-1C-F c documented an excellent in vivo stability and a high tissue bioavailability. As effective and inexpensive drug candidates, Nanosota-1 may contribute to the battle against COVID-19.
- Department of Veterinary and Biomedical Sciences, University of Minnesota, Saint Paul, United States.
Organizational Affiliation: