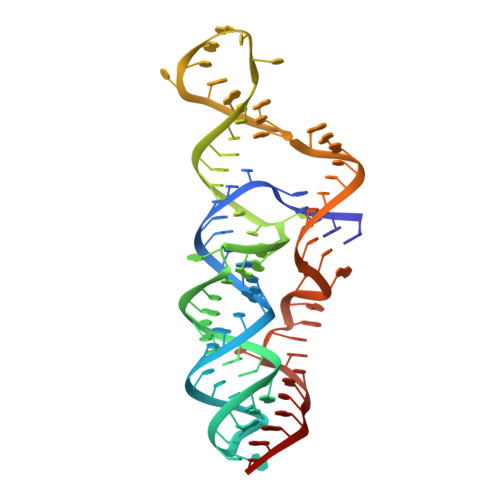
Three-dimensional structure of a flavivirus dumbbell RNA reveals molecular details of an RNA regulator of replication.
Akiyama, B.M., Graham, M.E., O Donoghue, Z., Beckham, J.D., Kieft, J.S.(2021) Nucleic Acids Res 49: 7122-7138
- PubMed: 34133732 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkab462
- Primary Citation Related Structures:
7KGA - PubMed Abstract:
Mosquito-borne flaviviruses (MBFVs) including dengue, West Nile, yellow fever, and Zika viruses have an RNA genome encoding one open reading frame flanked by 5' and 3' untranslated regions (UTRs). The 3' UTRs of MBFVs contain regions of high sequence conservation in structured RNA elements known as dumbbells (DBs). DBs regulate translation and replication of the viral RNA genome, functions proposed to depend on the formation of an RNA pseudoknot. To understand how DB structure provides this function, we solved the x-ray crystal structure of the Donggang virus DB to 2.1Å resolution and used structural modeling to reveal the details of its three-dimensional fold. The structure confirmed the predicted pseudoknot and molecular modeling revealed how conserved sequences form a four-way junction that appears to stabilize the pseudoknot. Single-molecule FRET suggests that the DB pseudoknot is a stable element that can regulate the switch between translation and replication during the viral lifecycle by modulating long-range RNA conformational changes.
- Department of Biochemistry and Molecular Genetics, Aurora, CO 80045, USA.
Organizational Affiliation: