Molecular basis for a germline-biased neutralizing antibody response to SARS-CoV-2.
Clark, S.A., Clark, L.E., Pan, J., Coscia, A., McKay, L.G.A., Shankar, S., Johnson, R.I., Griffiths, A., Abraham, J.(2020) bioRxiv
- PubMed: 33200128 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1101/2020.11.13.381533
- Primary Citation Related Structures:
7KFV, 7KFW, 7KFX, 7KFY - PubMed Abstract:
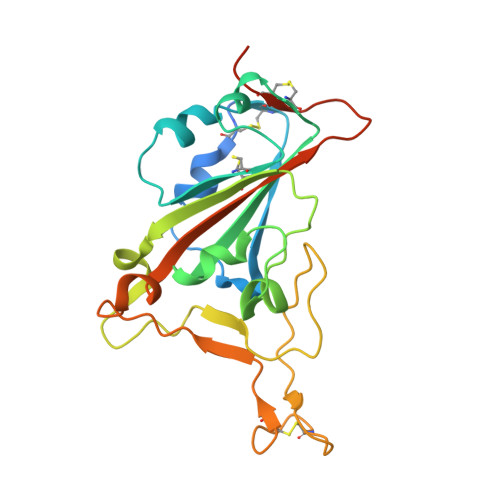
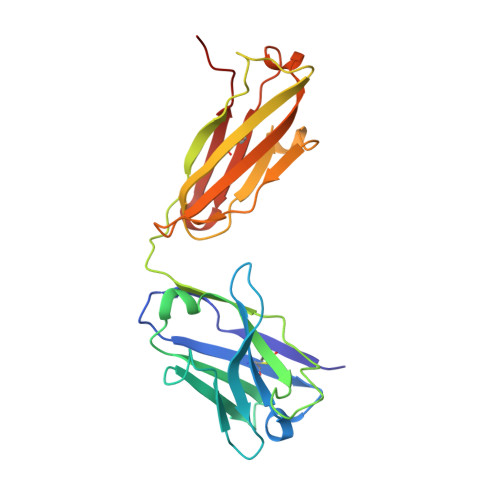
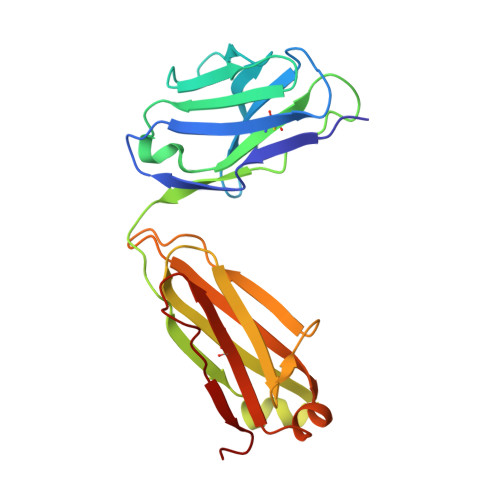
The SARS-CoV-2 viral spike (S) protein mediates attachment and entry into host cells and is a major target of vaccine and drug design. Potent SARS-CoV-2 neutralizing antibodies derived from closely related antibody heavy chain genes (IGHV3-53 or 3-66) have been isolated from multiple COVID-19 convalescent individuals. These usually contain minimal somatic mutations and bind the S receptor-binding domain (RBD) to interfere with attachment to the cellular receptor angiotensin-converting enzyme 2 (ACE2). We used antigen-specific single B cell sorting to isolate S-reactive monoclonal antibodies from the blood of a COVID-19 convalescent individual. The seven most potent neutralizing antibodies were somatic variants of the same IGHV3-53-derived antibody and bind the RBD with varying affinity. We report X-ray crystal structures of four Fab variants bound to the RBD and use the structures to explain the basis for changes in RBD affinity. We show that a germline revertant antibody binds tightly to the SARS-CoV-2 RBD and neutralizes virus, and that gains in affinity for the RBD do not necessarily correlate with increased neutralization potency, suggesting that somatic mutation is not required to exert robust antiviral effect. Our studies clarify the molecular basis for a heavily germline-biased human antibody response to SARS-CoV-2.