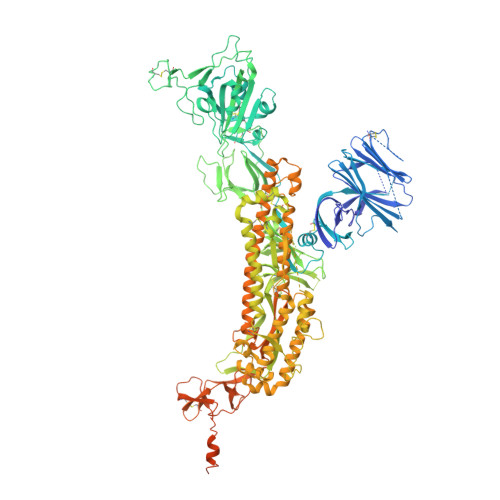
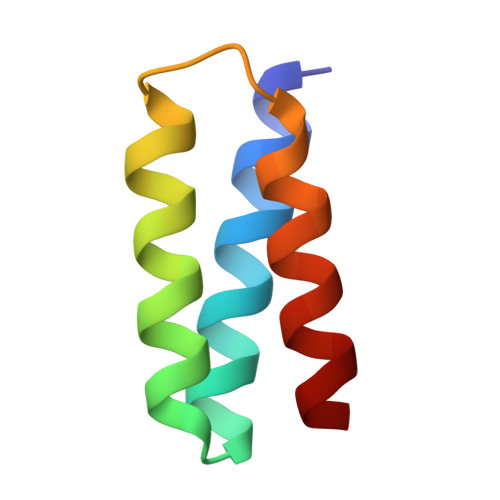
De novo design of picomolar SARS-CoV-2 miniprotein inhibitors.
Cao, L., Goreshnik, I., Coventry, B., Case, J.B., Miller, L., Kozodoy, L., Chen, R.E., Carter, L., Walls, A.C., Park, Y.J., Strauch, E.M., Stewart, L., Diamond, M.S., Veesler, D., Baker, D.(2020) Science 370: 426-431
- PubMed: 32907861 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/science.abd9909
- Primary Citation Related Structures:
7JZL, 7JZM, 7JZN, 7JZU - PubMed Abstract:
Targeting the interaction between the severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) spike protein and the human angiotensin-converting enzyme 2 (ACE2) receptor is a promising therapeutic strategy. We designed inhibitors using two de novo design approaches. Computer-generated scaffolds were either built around an ACE2 helix that interacts with the spike receptor binding domain (RBD) or docked against the RBD to identify new binding modes, and their amino acid sequences were designed to optimize target binding, folding, and stability. Ten designs bound the RBD, with affinities ranging from 100 picomolar to 10 nanomolar, and blocked SARS-CoV-2 infection of Vero E6 cells with median inhibitory concentration (IC 50 ) values between 24 picomolar and 35 nanomolar. The most potent, with new binding modes, are 56- and 64-residue proteins (IC 50 ~ 0.16 nanograms per milliliter). Cryo-electron microscopy structures of these minibinders in complex with the SARS-CoV-2 spike ectodomain trimer with all three RBDs bound are nearly identical to the computational models. These hyperstable minibinders provide starting points for SARS-CoV-2 therapeutics.
- Department of Biochemistry, University of Washington, Seattle, WA 98195, USA.
Organizational Affiliation: