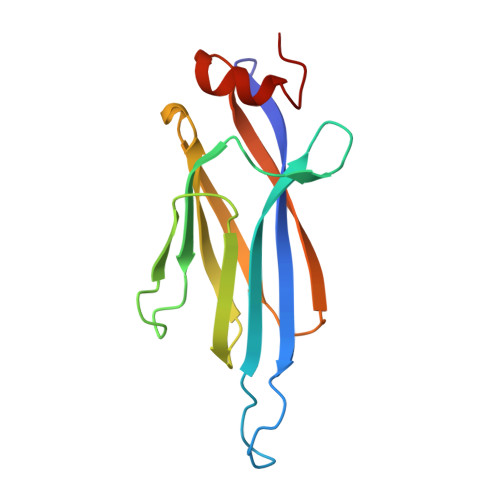
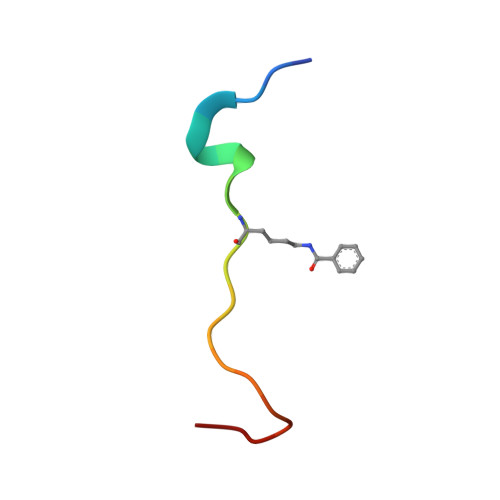
Global profiling of regulatory elements in the histone benzoylation pathway.
Wang, D., Yan, F., Wu, P., Ge, K., Li, M., Li, T., Gao, Y., Peng, C., Chen, Y.(2022) Nat Commun 13: 1369-1369
- PubMed: 35296687 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-022-29057-2
- Primary Citation Related Structures:
7F3S, 7F4A, 7F4E, 7F51, 7F5M - PubMed Abstract:
Lysine benzoylation (Kbz) is a recently discovered post-translational modification associated with active transcription. However, the proteins for maintaining and interpreting Kbz and the physiological roles of Kbz remain elusive. Here, we systematically characterize writer, eraser, and reader proteins of histone Kbz in S. cerevisiae using proteomic, biochemical, and structural approaches. Our study identifies 27 Kbz sites on yeast histones that can be regulated by cellular metabolic states. The Spt-Ada-Gcn5 acetyltransferase (SAGA) complex and NAD + -dependent histone deacetylase Hst2 could function as the writer and eraser of histone Kbz, respectively. Crystal structures of Hst2 complexes reveal the molecular basis for Kbz recognition and catalysis by Hst2. In addition, we demonstrate that a subset of YEATS domains and bromodomains serve as Kbz readers, and structural analyses reveal how YEATS and bromodomains recognize Kbz marks. Moreover, the proteome-wide screening of Kbz-modified proteins identifies 207 Kbz sites on 149 non-histone proteins enriched in ribosome biogenesis, glycolysis/gluconeogenesis, and rRNA processing pathways. Our studies identify regulatory elements for the Kbz pathway and provide a framework for dissecting the biological functions of lysine benzoylation.
- State Key Laboratory of Molecular Biology, Shanghai Institute of Biochemistry and Cell Biology, Center for Excellence in Molecular Cell Science, Chinese Academy of Sciences, 200031, Shanghai, China.
Organizational Affiliation: