Structural mechanism of calcium-mediated hormone recognition and G beta interaction by the human melanocortin-1 receptor.
Ma, S., Chen, Y., Dai, A., Yin, W., Guo, J., Yang, D., Zhou, F., Jiang, Y., Wang, M.W., Xu, H.E.(2021) Cell Res 31: 1061-1071
- PubMed: 34453129 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41422-021-00557-y
- Primary Citation Related Structures:
7F4D, 7F4F, 7F4H, 7F4I - PubMed Abstract:
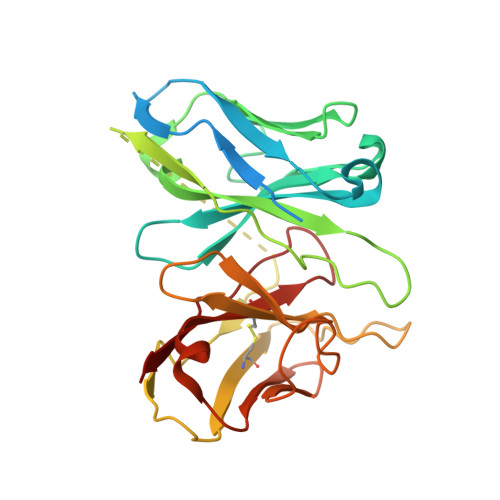
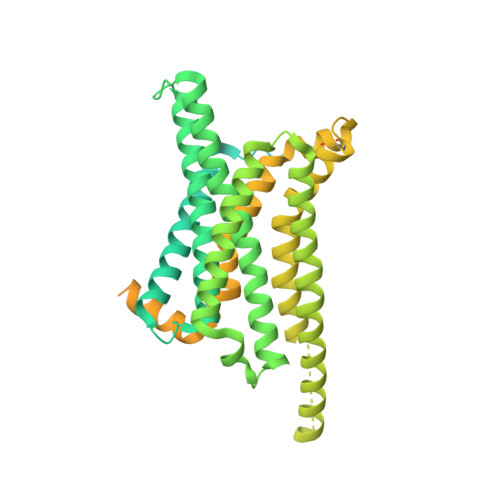
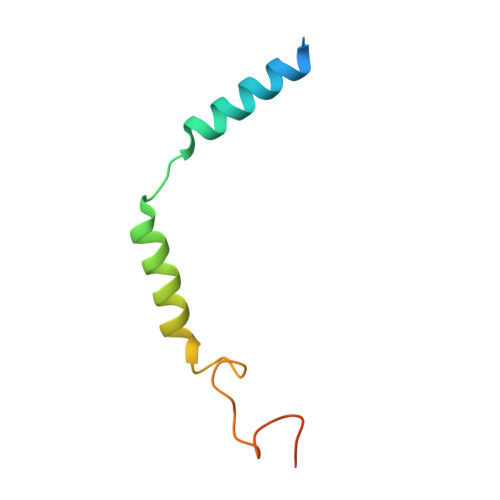
Melanocortins are peptide hormones critical for the regulation of stress response, energy homeostasis, inflammation, and skin pigmentation. Their functions are mediated by five G protein-coupled receptors (MC1R-MC5R), predominately through the stimulatory G protein (Gs). MC1R, the founding member of melanocortin receptors, is mainly expressed in melanocytes and is involved in melanogenesis. Dysfunction of MC1R is associated with the development of melanoma and skin cancer. Here we present three cryo-electron microscopy structures of the MC1R-Gs complexes bound to endogenous hormone α-MSH, a marketed drug afamelanotide, and a synthetic agonist SHU9119. These structures reveal the orthosteric binding pocket for the conserved HFRW motif among melanocortins and the crucial role of calcium ion in ligand binding. They also demonstrate the basis of differential activities among different ligands. In addition, unexpected interactions between MC1R and the Gβ subunit were discovered from these structures. Together, our results elucidate a conserved mechanism of calcium-mediated ligand recognition, a specific mode of G protein coupling, and a universal activation pathway of melanocortin receptors.
- The CAS Key Laboratory of Receptor Research, Shanghai Institute of Materia Medica, Chinese Academy of Sciences, Shanghai, China.
Organizational Affiliation: