Novel sarbecovirus bispecific neutralizing antibodies with exceptional breadth and potency against currently circulating SARS-CoV-2 variants and sarbecoviruses.
Wang, Y., Liu, M., Shen, Y., Ma, Y., Li, X., Zhang, Y., Liu, M., Yang, X.L., Chen, J., Yan, R., Luan, D., Wang, Y., Chen, Y., Wang, Q., Lin, H., Li, Y., Wu, K., Zhu, T., Zhao, J., Lu, H., Wen, Y., Jiang, S., Wu, F., Zhou, Q., Shi, Z.L., Huang, J.(2022) Cell Discov 8: 36-36
- PubMed: 35443747 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41421-022-00401-6
- Primary Citation Related Structures:
7EPX - PubMed Abstract:
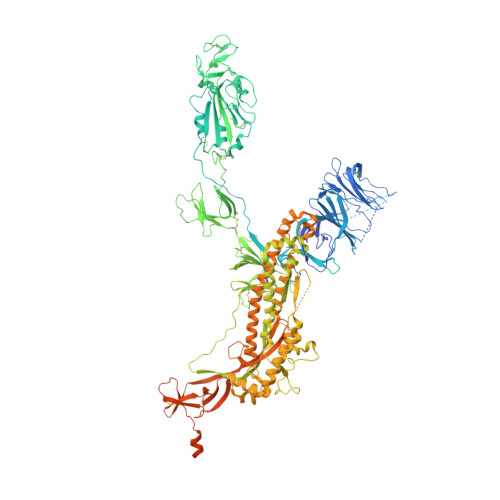
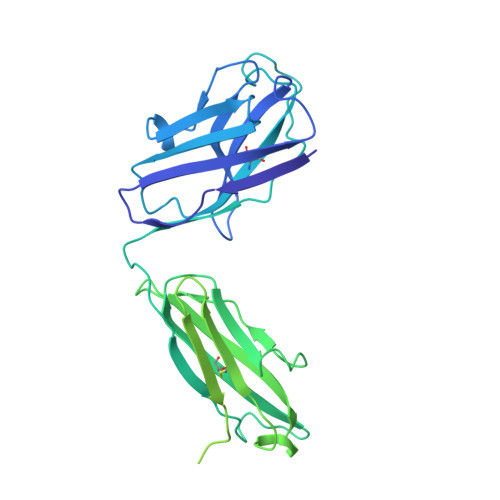
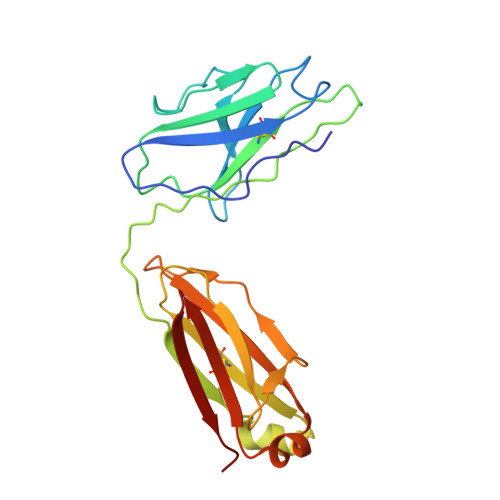
The emergence of severe acute respiratory syndrome coronavirus-2 (SARS-CoV-2) variants of concern, including Alpha (B.1.1.7), Beta (B.1.351), Gamma (P.1), Delta (B.1.617.2), and Omicron (B.1.1.529) has aroused concerns over their increased infectivity and transmissibility, as well as decreased sensitivity to SARS-CoV-2-neutralizing antibodies (NAbs) and the current coronavirus disease 2019 (COVID-19) vaccines. Such exigencies call for the development of pan-sarbecovirus vaccines or inhibitors to combat the circulating SARS-CoV-2 NAb-escape variants and other sarbecoviruses. In this study, we isolated a broadly NAb against sarbecoviruses named GW01 from a donor who recovered from COVID-19. Cryo-EM structure and competition assay revealed that GW01 targets a highly conserved epitope in a wide spectrum of different sarbecoviruses. However, we found that GW01, the well-known sarbecovirus NAb S309, and the potent SARS-CoV-2 NAbs CC12.1 and REGN10989 only neutralize about 90% of the 56 tested currently circulating variants of SARS-CoV-2 including Omicron. Therefore, to improve efficacy, we engineered an IgG-like bispecific antibody GW01-REGN10989 (G9) consisting of single-chain antibody fragments (scFv) of GW01 and REGN10989. We found that G9 could neutralize 100% of NAb-escape mutants (23 out of 23), including Omicron variant, with a geometric mean (GM) 50% inhibitory concentration of 8.8 ng/mL. G9 showed prophylactic and therapeutic effects against SARS-CoV-2 infection of both the lung and brain in hACE2-transgenic mice. Site-directed mutagenesis analyses revealed that GW01 and REGN10989 bind to the receptor-binding domain in different epitopes and from different directions. Since G9 targets the epitopes for both GW01 and REGN10989, it was effective against variants with resistance to GW01 or REGN10989 alone and other NAb-escape variants. Therefore, this novel bispecific antibody, G9, is a strong candidate for the treatment and prevention of infection by SARS-CoV-2, NAb-escape variants, and other sarbecoviruses that may cause future emerging or re-emerging coronavirus diseases.
- Key Laboratory of Medical Molecular Virology (MOE/NHC/CAMS) and Shanghai Institute of Infectious Disease and Biosecurity, Shanghai Public Health Clinical Center, School of Basic Medical Sciences, Fudan University, Shanghai, China.
Organizational Affiliation: