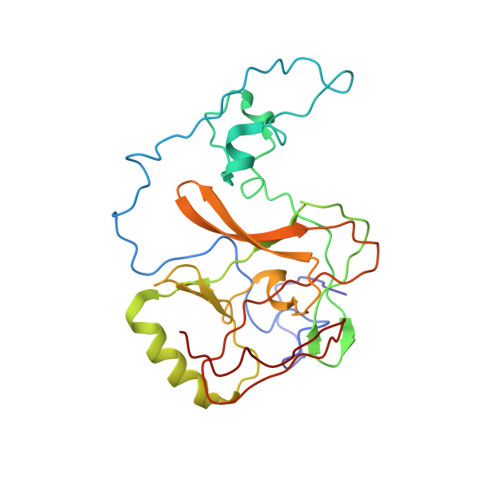
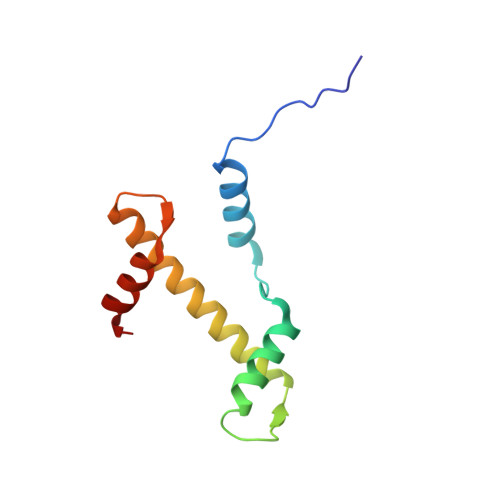
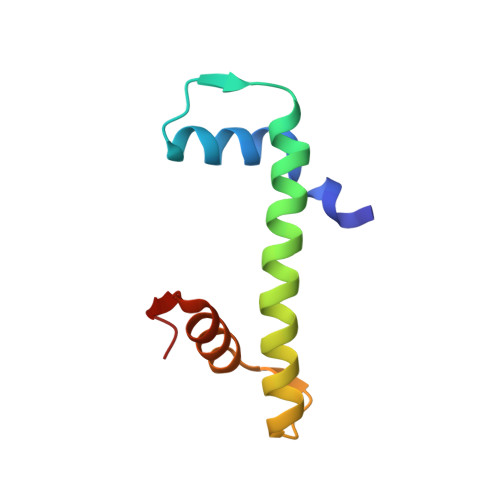
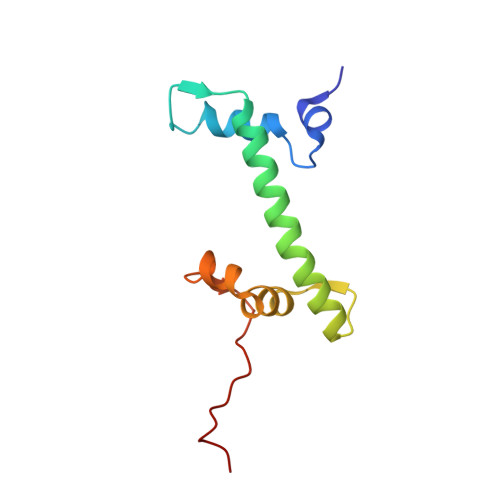
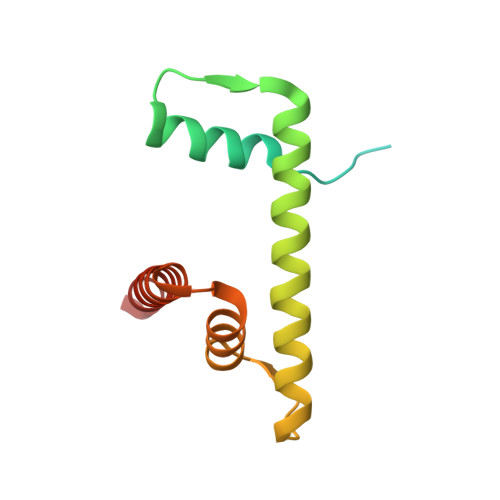
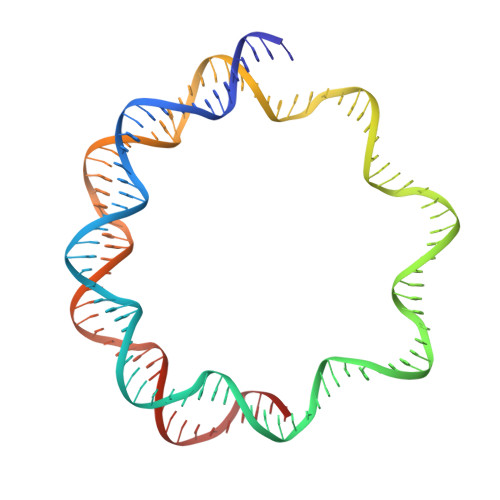
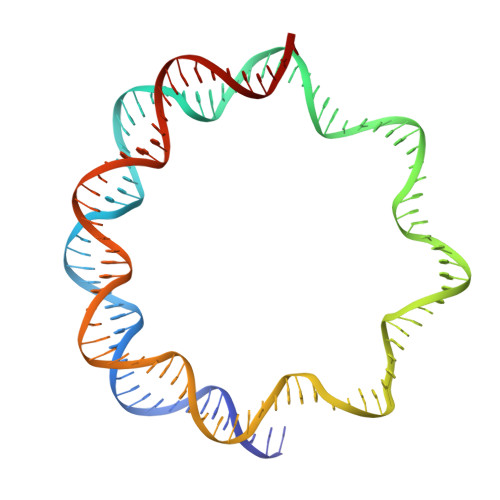
Cryo-EM structure of SETD2/Set2 methyltransferase bound to a nucleosome containing oncohistone mutations.
Liu, Y., Zhang, Y., Xue, H., Cao, M., Bai, G., Mu, Z., Yao, Y., Sun, S., Fang, D., Huang, J.(2021) Cell Discov 7: 32-32
- PubMed: 33972509 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41421-021-00261-6
- Primary Citation Related Structures:
7EA5, 7EA8 - PubMed Abstract:
Substitution of lysine 36 with methionine in histone H3.3 (H3.3K36M) is an oncogenic mutation that inhibits SETD2-mediated histone H3K36 tri-methylation in tumors. To investigate how the oncohistone mutation affects the function of SETD2 at the nucleosome level, we determined the cryo-EM structure of human SETD2 associated with an H3.3K36M nucleosome and cofactor S-adenosylmethionine (SAM), and revealed that SETD2 is attached to the N-terminal region of histone H3 and the nucleosome DNA at superhelix location 1, accompanied with the partial unwrapping of nucleosome DNA to expose the SETD2-binding site. These structural features were also observed in the previous cryo-EM structure of the fungal Set2-nucleosome complex. By contrast with the stable association of SETD2 with the H3.3K36M nucleosome, the EM densities of SETD2 could not be observed on the wild-type nucleosome surface, suggesting that the association of SETD2 with wild-type nucleosome might be transient. The linker histone H1, which stabilizes the wrapping of nucleosome DNA at the entry/exit sites, exhibits an inhibitory effect on the activities of SETD2 and displays inversely correlated genome distributions with that of the H3K36me3 marks. Cryo-EM analysis of yeast H3K36 methyltransferase Set2 complexed with nucleosomes further revealed evolutionarily conserved structural features for nucleosome recognition in eukaryotes, and provides insights into the mechanism of activity regulation. These findings have advanced our understanding of the structural basis for the tumorigenesis mechanism of the H3.3K36M mutation and highlight the effect of nucleosome conformation on the regulation of histone modification.
- Ninth People's Hospital, Shanghai Jiao Tong University School of Medicine, Shanghai, 200125, China.
Organizational Affiliation: