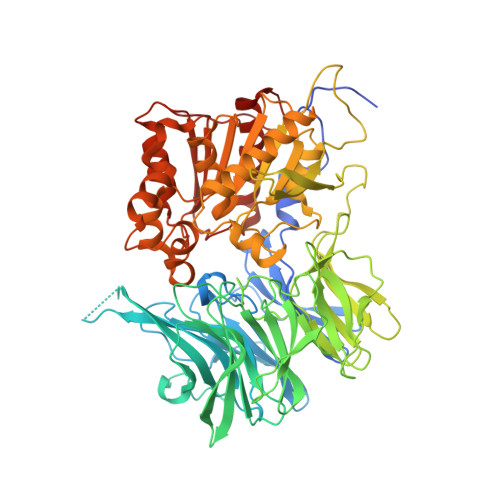
Mycobacterium tuberculosis puromycin hydrolase displays a prolyl oligopeptidase fold and an acyl aminopeptidase activity.
Zhao, Y., Feng, Q., Zhou, X., Zhang, Y., Lukman, M., Jiang, J., Ruiz-Carrillo, D.(2021) Proteins 89: 614-622
- PubMed: 33426726 Search on PubMed
- DOI: https://doi.org/10.1002/prot.26044
- Primary Citation Related Structures:
7C72 - PubMed Abstract:
Puromycin-hydrolizing peptidases have been described as members of the prolyl oligopeptidase peptidase family. These enzymes are present across all domains of life but still little is known of the homologs found in the pathogenic bacterium Mycobacterium tuberculosis. The crystal structure of a M. tuberculosis puromycin hydrolase peptidase has been determined at 3 Angstrom resolution, revealing a conserved prolyl oligopeptidase fold, defined by α/β-hydrolase and β-propeller domains with two distinctive loops that occlude access of large substrates to the active site. The enzyme displayed amino peptidase activity with a substrate specificity preference for hydrophobic residues in the decreasing order of phenylalanine, leucine, alanine and proline. The enzyme's active site is lined by residues Glu564 for the coordination of the substrates amino terminal moiety and His561, Val608, Tyr78, Trp306, Phe563 and Ty567 for the accommodation of hydrophobic substrates. The availability of a crystal structure for puromycin hydrolase of M. tuberculosis shall facilitate the development of inhibitors with therapeutic applications.
- Department of Biological Sciences, School of Science, Xi'an Jiaotong-Liverpool University, Suzhou, Jiangsu, China.
Organizational Affiliation: