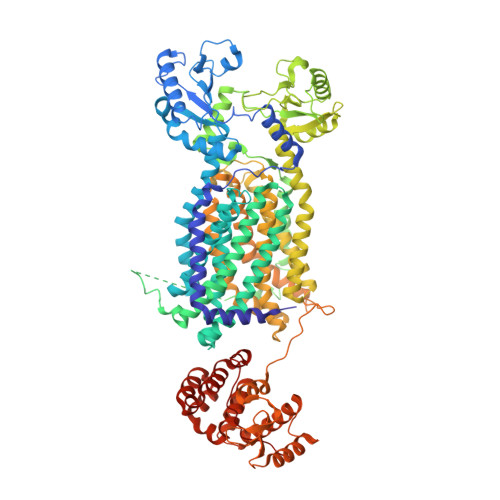
Structural Basis for the Inhibition of Mycobacterial MmpL3 by NITD-349 and SPIRO.
Yang, X., Hu, T., Yang, X., Xu, W., Yang, H., Guddat, L.W., Zhang, B., Rao, Z.(2020) J Mol Biology 432: 4426-4434
- PubMed: 32512002 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2020.05.019
- Primary Citation Related Structures:
7C2M, 7C2N - PubMed Abstract:
Novel antitubercular agents are urgently needed to combat the emergence of global drug resistance to human tuberculosis. Mycobacterial membrane protein Large 3 (MmpL3) is a promising drug target because its activity is essential and required for cell-wall biosynthesis. Several classes of MmpL3 inhibitors have been developed against Mycobacterium tuberculosis (Mtb) with potent anti-tuberculosis activity. These include the drug candidate SQ109, which has progressed to phase IIb/III clinical trials. Here, we have determined the crystal structures of MmpL3 in complex with NITD-349 and SPIRO. Both inhibitors bind deep in the central channel of transmembrane domain and cause conformational changes to the protein. The amide nitrogen and indole nitrogen of NITD-349 and the piperidine nitrogen of SPIRO interact and clamp Asp645. Structural analysis of the two structures reveals that these inhibitors target the proton relay pathway to block the activity of MmpL3. The findings presented here enrich our understanding of the binding modes of MmpL3 inhibitors and provide directions to enable further rational drug design targeting MmpL3.
- Shanghai Institute for Advanced Immunochemical Studies and School of Life Science and Technology, ShanghaiTech University, Shanghai 201210, China; CAS Center for Excellence in Molecular Cell Science, Shanghai Institute of Biochemistry and Cell Biology, Chinese Academy of Sciences, Shanghai 200031, China; University of Chinese Academy of Sciences, Beijing 100101, China.
Organizational Affiliation: