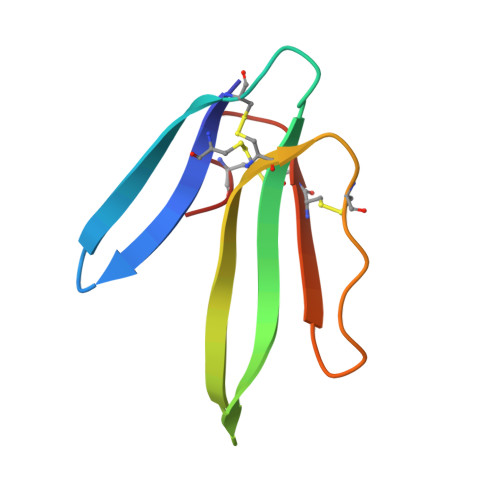
Unusual quaternary structure of a homodimeric synergistic-type toxin from mamba snake venom defines its molecular evolution.
Aoki-Shioi, N., Jobichen, C., Sivaraman, J., Kini, R.M.(2020) Biochem J 477: 3951-3962
- PubMed: 33000863 Search on PubMed
- DOI: https://doi.org/10.1042/BCJ20200529
- Primary Citation Related Structures:
7C28 - PubMed Abstract:
Snake venoms are complex mixtures of enzymes and nonenzymatic proteins that have evolved to immobilize and kill prey animals or deter predators. Among them, three-finger toxins (3FTxs) belong to the largest superfamily of nonenzymatic proteins. They share a common structure of three β-stranded loops extending like fingers from a central core containing all four conserved disulfide bonds. Most 3FTxs are monomers and through subtle changes in their amino acid sequences, they interact with different receptors, ion channels and enzymes to exhibit a wide variety of biological effects. The 3FTxs have further expanded their pharmacological space through covalent or noncovalent dimerization. Synergistic-type toxins (SynTxs) isolated from the deadly mamba venoms, although nontoxic, have been known to enhance the toxicity of other venom proteins. However, the details of three-dimensional structure and molecular mechanism of activity of this unusual class of 3FTxs are unclear. We determined the first three-dimensional structure of a SynTx isolated from Dendroaspis jamesoni jamesoni (Jameson's mamba) venom. The SynTx forms a unique homodimer that is held together by an interchain disulfide bond. The dimeric interface is elaborate and encompasses loops II and III. In addition to the inter-subunit disulfide bond, the hydrogen bonds and hydrophobic interactions between the monomers contribute to the dimer formation. Besides, two sulfate ions that mediate interactions between the monomers. This unique quaternary structure is evolved through noncovalent homodimers such as κ-bungarotoxins. This novel dimerization further enhances the diversity in structure and function of 3FTxs.
- Department of Chemistry, Faculty of Science, Fukuoka University, 19-1, 8-chome Nanakuma, Jonan-ku, Fukuoka 814-0180, Japan.
Organizational Affiliation: